Jeremy Leung
Century of the gene
In October 1990, biologists officially embarked on one of the century’s most ambitious scientific efforts: reading the 3 billion pairs of genetic subunits — the A’s, T’s, C’s and G’s — that make up the human instruction book. The project has been compared to the Manhattan Project to build an atomic bomb and the Apollo missions to land on the moon. The effort promised to blow open our understanding of basic biology, reveal relationships between the myriad forms of life on the planet and transform medicine through insights into genetic diseases and potential cures. When a draft of the instruction book was announced in 2000, the scientists having read essentially every letter, President Bill Clinton called it a “stunning and humbling achievement” and predicted that it would “revolutionize the diagnosis, prevention and treatment of most, if not all, human diseases.”

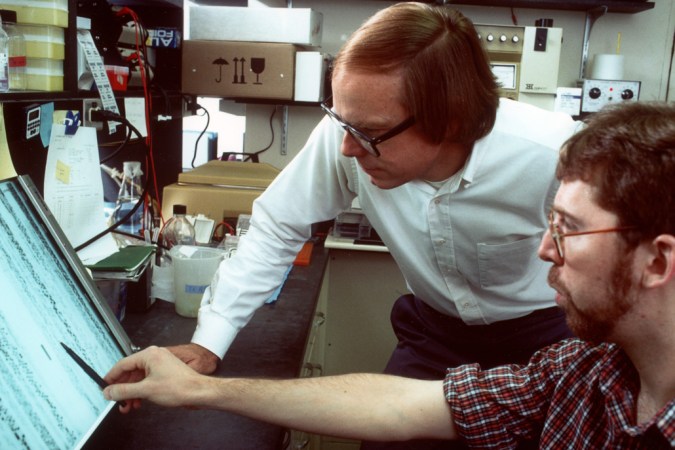
Even dreaming up such an endeavor depended on decades of previous discoveries. Most of our textbook notions of how the information of life is encoded, the details of how it’s passed on and how it makes us who we are, were uncovered during the 20th century. In 1905, English biologist William Bateson, who championed the work of Austrian monk Gregor Mendel, suggested the term genetics for a new field of study focused on heredity and variation. Early the next decade, American biologist Thomas Hunt Morgan and his colleagues showed that genes were carried on chromosomes. Biochemists had been studying DNA for nearly three-quarters of a century when Oswald Avery and his team at the Rockefeller Institute in New York City helped establish in the 1940s that DNA is the genetic material. And perhaps most notable, and famous today, is the 1953 discovery of the double-helix structure of DNA, by James Watson and Francis Crick of University of Cambridge and Rosalind Franklin and Maurice Wilkins of King’s College London.

But when the draft instruction book was published, independently by an international collective of academic and government labs called the Human Genome Project and the private company Celera Genomics, led by J. Craig Venter, the text was “as striking for what we don’t see as for what we do,” Science News reported. There were many fewer genes than expected, leaving a puzzle about what all the remaining DNA was for. In the decades since, scientists have filled in some of that puzzle — identifying hosts of genes, for example, that don’t make proteins but are still essential in the body. Other researchers have searched the instruction book to find new treatments for diseases and to figure out how we’re all related — not just people, but all life on planet Earth, past and present.

To explore how far our understanding of our DNA has come, Science News senior writer and molecular biology reporter Tina Hesman Saey talked with Eric Green, director of the National Human Genome Research Institute at the National Institutes of Health in Bethesda, Md. Green got his start in genomics in the lab of Maynard Olson at Washington University in St. Louis, a pioneer in the field. At the same time, Saey was a graduate student in molecular genetics working down the hall. She remembers as an undergraduate student sequencing the genes of bacteria 50 to 100 chemical subunits, or bases, at a time. “My mind was completely blown by the idea that you could put together 3 billion bases.” The conversation that follows, which has been edited for length and clarity, looks back on the project and ahead to all that’s left to learn.
— Elizabeth Quill
An ambitious undertaking

Saey: My first memory of the Human Genome Project was when I was an undergraduate student at the University of Nebraska in Lincoln, and I remember Walter Gilbert, who is a Nobel Prize winner, coming and talking about the project. He proposed this really audacious thing of sequencing 3 billion pairs of bases in the human genome — all of our DNA. After Gilbert’s talk, I walked back to the lab with a couple of professors, and they were saying, ‘This can never happen. It’s going to cost way too much money. There’s just no way we can do this.’ So how did you pull it off?
Green: By the time the genome project started in October of 1990, I was working in a real cutting-edge genomics lab at Washington University. We were actually one of the first funded groups to participate in the Human Genome Project. We had some ideas on how to start, and we had really no idea how we were going to pull it off.
It was the overwhelmingly compelling vision for why this was so important that galvanized enthusiasm among not only a group of scientists like myself, but also the funding agencies, the governments, the private funders from around the world, who said, “This seems unimaginable, like putting a person on the moon, but it seems so important. We’ll figure it out.” So it was one of these circumstances where you just get the right people in the right place, get them resourced, get them organized, be willing to do things differently, and then figure it out as you go.

Saey: I got to witness this because I was a graduate student at Washington University in St. Louis, in a lab sequencing the yeast genome. And Robert Waterston’s lab, which received one of the first grants from the Human Genome Project, was right across the hall. They started out doing the C. elegans, the roundworm genome. I remember they were starting very methodically, mapping out the genes and then sequencing each piece, marching along. But then, towards the end of the ’90s, there was this shotgun sequencing revolution. That was spearheaded by kind of a controversial figure, Craig Venter. You just shred the genome, throw it all in a sequencing machine and then put it together in the computer. Did that help a lot?
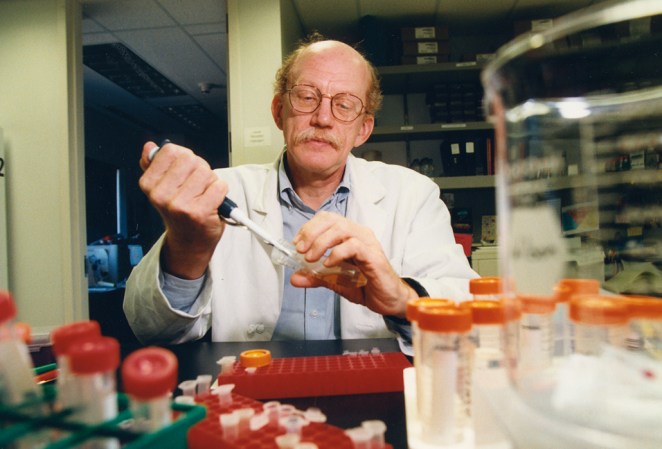

Green: There’s no question it sped things up. What Craig successfully did was to determine that there were approaches that could be used where you didn’t have to do piecemeal sequencing. The important nuance to point out is the only way you’re able to put [the pieces] back together then was by having many mapping elements that allow you to hang pieces together and organize them. It’s not like it all zipped together 3 billion letters. A lot of the meticulous mapping that had been done, painstaking mapping, helped provide organizing guideposts.
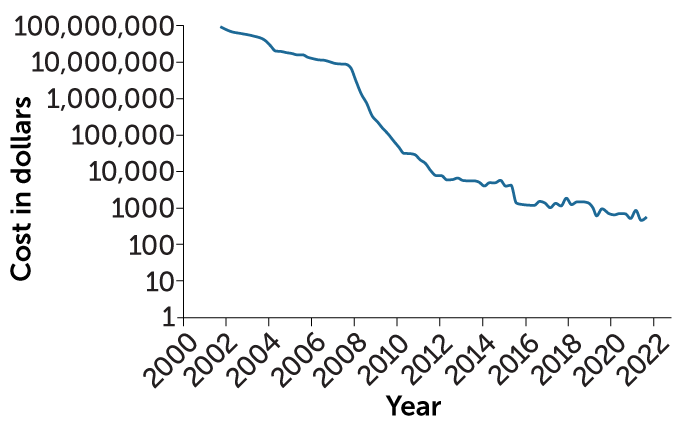
The press covered it as a race, and the press covered it as option A versus option B. And the truth resided somewhere in between. What was driving the change, of course, was technology advances. If you chart the time since the end of the Human Genome Project, it’s the same phenomenon. Every single time there’s a technology surge, you find yourself doing things completely different than the way you used to. In fact, you almost laugh about it.
Saey: Technology has come a very long way from what I was doing. It’s moved into things I can’t even imagine. You can sequence thousands of bases at a time now.
Green: The other part of the story that sometimes doesn’t get told: It’s not even just the laboratory bench-based technologies. It’s also the computational technologies. Some people don’t realize that when the Human Genome Project started, there was not really a widely functional internet. I was just barely starting to use e-mail.
So here it was, we were one of the first funded groups for the Human Genome Project. We were considered the cutting-edge, state of the art. We were collaborating with an outside group generating some sequences, and the only practical way for my collaborator to get me the 300 to 400 bases of sequence was to handwrite it on a piece of paper and fax it to me. And I would analyze it by eye. It’s just remarkable that that was where we were when the genome project started.
Big surprises
Saey: In 2000 was the big press conference to announce the rough draft of the human genome. I was just starting my journalism career at the St. Louis Post-Dispatch and reported on this. At that time, it was a big revelation that it wasn’t just a bunch of genes strung out along the chromosome, that there were these big deserts in between genes, and that we didn’t have nearly as many genes as we thought we were going to. Humans are such complex organisms, how could we not have many more genes than a fruit fly, or a worm? That just didn’t make sense.
But now, I think, we are getting a better understanding largely because of the way we can analyze the genome. Can you talk about how that evolution in thinking has progressed? Even, what is a gene, for instance?
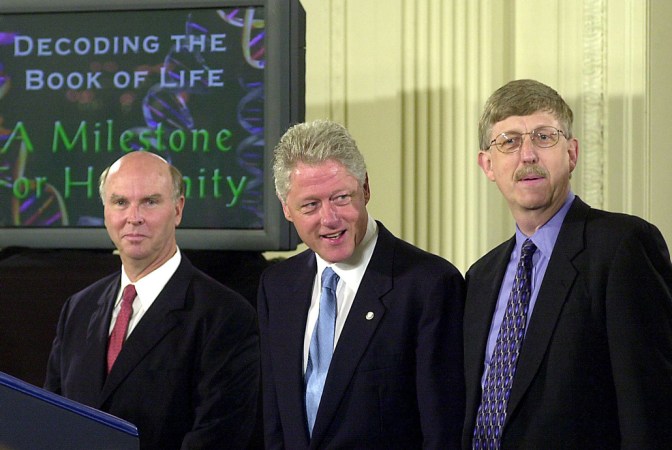

Green: Before the genome project started, some [people] were quite critical, and really said it was a bad idea. Some of their arguments was that it was a waste of time to sequence the genome end to end; we should just focus and sequence the genes, as if all of humans’ biological richness was going to reside in the genes. And we look back now, and thank goodness we didn’t listen to those critics. Because if we would have done the shortcut and only focused on the genes, we would have only skimmed the biological complexity of humans.
What we have come to learn is that while only 1.5 percent of the letters of the human genome directly encode for what are classically known as protein-coding genes — which is what we understand the best, DNA that gets made into RNA, which gets made into protein — there’s a much larger fraction of the human genome that is biologically important and evolutionarily conserved. It’s widened our definition of a gene, because we now know that sometimes DNA may make RNA and RNA may go off and do all sorts of biological things.
Then there’s a whole set of sequences that are far more plentiful than gene sequences, that are really doing all the choreography in our genomes in terms of determining when, where and how much genes get turned on, in what cells and what tissues, at what developmental stages, under what conditions, and so on and so forth.
It pushed us to think about all the other biological functions in DNA outside the genes. And as you accurately point out, we don’t really have a rulebook for that. And thank goodness the computer technology is helping us because the human eye would just fail miserably at figuring this out. And so as much as anything, computational biology, bioinformatics, data science are being used as the dominant research tools to help bring clarity as to how it is that noncoding sequences in the human genome function. And how they do that in a very carefully crafted choreography with the genes.
Saey: Well, I’m glad you brought up those sequences, because those are some of my favorites. Prior to becoming a journalist, I never really thought much about RNA, except that it’s kind of a pain to work with. But now, I’m a huge fan of noncoding RNAs [the RNAs that don’t go on to make proteins]. There are so many of them, there’s just such a huge variety of them. And they work in so many important ways.
I don’t think that 20 years ago we could have conceived that RNAs that didn’t make proteins would actually be important for something. The genes those RNAs were copied from were considered broken genes or pseudogenes. They were junk.
Green: Or sloppy transcription, that our enzymes are just going off and making a bunch of RNA because they don’t know how to control themselves. And it’s just garbage. But, no. And I like your point about 20 years ago, we couldn’t imagine. I would propose that 20 years from now, we might look back at this conversation and say, ‘Oh, my goodness, think about all these other ways that the genome functions.’ There’s no reason to think we have our hands around it all in terms of all the biological complexity of DNA; I’m quite sure we don’t.
Saey: And even when you find a protein-coding gene, you’re not just making one protein. You’re making, on average, seven or eight different versions of this protein from the same gene. After RNA gets copied from DNA, you can mix and match the little parts of a gene to make completely new proteins. And then you can tack on all of these other little chemical groups to change the way things work.
Green: When I was getting my Ph.D. at Washington University in the 1980s, I didn’t work on DNA, I didn’t work on molecular biology, I didn’t work on RNA. I was working on a set of proteins, studying how they had sugar molecules added to them after they were made, and how, depending upon what tissue they were made in, they got different structures of sugar molecules attached. So just as you point out, you start off with one gene, and you can end up with multiple RNAs that lead to multiple different proteins. And each of those proteins could have different post-translational modifications depending on what tissue, what conditions, what development stage, etcetera. This is the incredible amplification of complexity. It’s not in our gene number. We have a long way to go to fully understanding all this.
Saey: Another thing that really surprises people is how much of our genome is actually extinct viruses and transposons — transposons being these jumping genes that still hop around in our genome. Those transposons can occasionally cause problems, but we also got a lot of innovations from them, including the human placenta, and maybe some things about the way our brains work. So, we’re not even completely human. If you want to view it that way, we’re a lot virus.
Green: Right. We’re a lot virus. We’re also not all Homo sapiens. Many, many people carry Neandertal DNA bits from a time when Neandertals and Homo sapiens both coexisted, and actually interbred. But not everybody in the world has that, which is also interesting. One of the aspects of genomics is that it not only has taught us and given us the biological instruction book, it’s also given us a fascinating record of evolution. We can use it to learn lots of things about our evolution, about human migrations, about aspects of humans on this globe.
Saey: How important is data that we get from sequencing the genomes of animals?
Green: The field of comparative genomics is one I used to work in, and one that I still think is incredibly exciting. It’s basically the idea of comparing our genome, especially the sequence level, to the genomes of other animals, at different evolutionary points. It’s like opening up evolution’s notebooks for every one of these critters. And there’s always things that you can learn by reviewing what evolution did and did not tolerate in terms of changes in the DNA. Evolution is a very powerful force, and it took notes. So it’s been a very valuable tool for teaching us what evolution was thinking, which in many ways is driven by biological innovation, biological function, and that has given us some clues about what’s really important in the human genome.
Focus on diversity
Saey: Most people who are interacting with DNA and with the genome these days do it through ancestry testing and consumer DNA testing. So you can identify the part of the world that people’s DNA came from. And that gets into a lot of discussion about race, and whether that has a biological basis, and what that might mean for medicine.
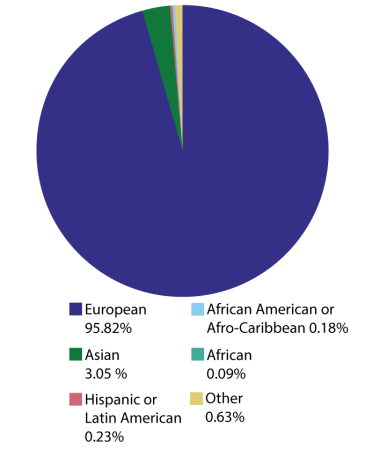
There’s been a lot of criticism lately of genetics and genomics, because it’s based a lot on the DNA of people of European ancestry — white people like you and me. But there’s a huge amount of genetic diversity in the world among humans, and especially in Africa, where humans got started. So what are we doing about getting a handle on the vast array of diversity that humans have?
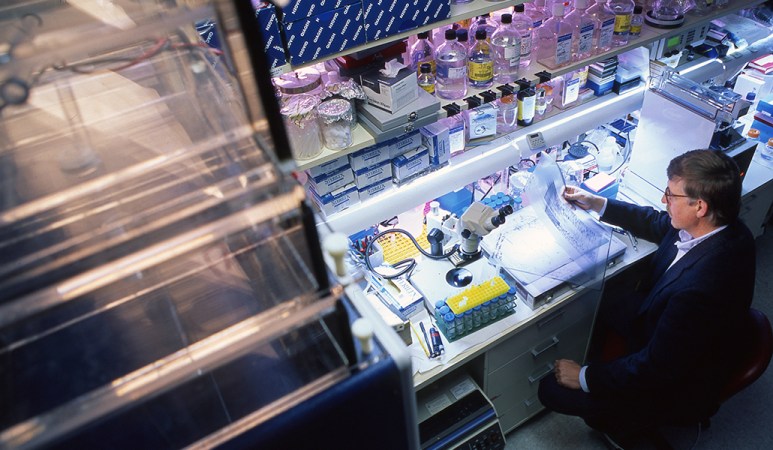

Green: There’s no question that the successes in genomics that we’ve been discussing are worth talking about and worth showcasing. At the same time, I’m very honest in saying that, as a field, we have not been perfect. One of the things that we just have to admit that we’ve really not been as successful on is making sure we’ve captured enough of the diversity of the human population with respect to the samples that we’ve used for doing genetic and genomic studies. We have got to catch up, we’ve got to fix this problem. It’s a very high priority.
I really want to emphasize, it’s not even just that it’s the socially right thing to do, that everybody should have information about their genomes. This is very important medically. We will make important inferences based on knowledge about variants that exist in genomes, we will do that through studies where we really need to have matching populations. And if the only populations we have a lot of genomic data on are people of European descent, we limit our ability to move genomic analyses and eventually genomic medicine into populations that are not of European descent. And so there’s a high priority through a number of efforts around the world, but including in the U.S., to work hard to capture much more diversity of the world’s populations in all studies that we do.
Saey: There’s been a lot of talk about racialized medicine, where you might have a person come in who is African American, and then you would say, ‘Oh, well, we should consider this to be the genome that we look at.’ Is that a good approach to take? Or do you think it should be broader somehow?
Green: The truth is, of course, there are certain diseases that tend to cluster in certain populations of common ancestry. And many times those are represented by racial groups. But racial grouping is really a social construct that has numerous imperfections. And so on the one hand, you can’t totally ignore some correlations that exist with certain diseases or certain responses to medications in certain groups. But it’s a very blunt tool to use. And we could do better. The way we could do this better is to track much more accurately to specific genomic features, as opposed to certain racial characteristics. So I think what we really want to pay attention to, and we will be doing this increasingly, is thinking about better ways of grouping and stratifying individuals and populations.
Saey: Well, I wanted to touch too on what we mean when we say genetic diversity. For the most part I think people are familiar with what scientists call SNPs, single nucleotide polymorphisms, and what other people might refer to as a mutation. But there’s lots of other ways that you can have diversity in the genome: You can be missing entire genes or entire chunks of chromosomes or you can have duplications of certain genes. Are we now able to look at that type of diversity as well? And do we know if that’s important?
Green: There’s no question that all forms of genomic diversity, genomic variation is probably the word I would use, are not only biologically relevant, they’re all proving to be medically relevant. Now, we don’t have a complete inventory of which ones are more relevant than others. But we already know of many examples where medically relevant variations in our genome can be a single letter, a string of letters, it could mean having extra letters or extra segments, or missing segments. It could be a rearrangement of segments. Every one of those [types of variations] are already known to be important in human disease, and eventually will be important for diagnostic medicine and the implementation of genomic medicine.
Saey: Do you envision a time when we will be able to study and interpret these bigger changes?
Green: I absolutely envision a time where people will get their complete genome sequenced end to end as part of their medical care, and maybe even at birth. I don’t think we’re there yet. But I truly believe that we will want that information as part of medical management. And I fully believe that technologies will become available and will be inexpensive enough to make it worthwhile. But those predictions are going to have to be based on evidence that indeed that’s feasible and valuable.
“I absolutely envision a time where people will get their complete genome sequenced end to end as part of their medical care, and maybe even at birth.”
Eric Green
Saey: I think most people thought that end-to-end sequencing is what we had in the early 2000s when we completed the Human Genome Project. But in fact, you weren’t really done at that point. Now you’re really done. Right? Can you talk a little bit about what it means to have a complete human genome sequence and what it took to get there?
Green: When the genome project ended, we attempted to be fairly transparent that we were declaring the genome project ending. We had generated the first reference sequence of the human genome to the best of our abilities that was essentially completed. By essentially, it was a great, great, great majority, well over 90 percent complete.
But we knew at that time that there were a number of places where we were simply incapable of reading the letters in the genome. Nearly all corresponded to regions of the human genome where the letters are very repetitive. The tricks that we use in the laboratory to read DNA, they just fail at that. We had no tools left in our toolbox at that time. It just seemed pointless to just say, ‘Well, we’ll just keep banging our head against the wall,’ even though we weren’t making progress. It’s just better to say, ‘Alright, we’re essentially done. And we will, over time, on a smaller scale, try to figure out how to fill in these gaps.’
In the last five years, you really saw the right technologies come onboard with the right people. The people working on the Human Genome Project were mostly biologists who learned computation. But this latest success came from people who were first trained in computation and then found out there was this big challenge in biology of getting the computer to help us put together these missing pieces using new technologies that allow you to take bigger, longer leaps across highly repetitive DNA.
Now they’ve generated end-to-end sequences of every human chromosome. There are major papers coming out. It will finally be a complete look at every letter of a human genome. And there might be some biologically important sequences that we were blind to until now.
What’s next?
Saey: So where do we go from here? If we’re done, what does the National Human Genome Research Institute do now?
Green: We recently finished a two-and-a-half-year strategic planning process to ask that very question for this coming decade. It was actually an overwhelming exercise because there were so many good ideas. We published these in Nature — our new 2020 strategic vision. Some of it [is] applications of genomics to medicine. Of course, everybody’s going to be excited about that. But there are many other forefronts of genomics that are just as exciting.
We still don’t have the perfect technologies that we can deploy anywhere in the world in any health setting, any medical study, that’ll get us end-to-end sequencing. We need better and cheaper technologies for letting us read human genome sequences inexpensively in clinical settings. We need complete end-to-end interpretation of every base of the human genome. We need to know not just about the genes, we need to know about all these noncoding regions. We need to understand every human variant that we can find in the world population. And we need to know is that variant biologically silent? Is it biologically relevant? Is it medically relevant? If it’s medically relevant, what’s the action that should be taken? That starts to point us to understanding the genomic basis of disease and also to think about how can we use information about genomic variation in the practice of medicine.
Also, [we need to] continue to think about what are the implications of these genomic advances to society? How are we going to make sure people understand this? How are we going to make sure things are applied equitably? How are we going to make sure it doesn’t exacerbate inequities in our society? How are we going to deal with a whole host of issues related to privacy?

Saey: I’m glad that you brought up equity and privacy, because those are some of the things that people are most concerned about right now. There are a lot of historically marginalized people who don’t want any part of genetic research because of the way their groups have been treated in the past. There’s been this history of colonialism. These groups say, if we’re going to do genetics on our people, then it should be our people doing it for us. What is NHGRI doing to build capacity in these communities so that they can do their own research and, maybe, if they decide they want to, share that with other people?
Green: I completely agree with the notion that if genomics is going to be a successful field, especially as we move this into medicine, we have got to make sure that we engage people from all different communities, all definitions of diversity, and make sure they benefit from it. We absolutely emphasize this point repeatedly in our 2020 strategic vision, so much so that the very first thing we did upon the new year in 2021 was to release what we call an action agenda for enhancing the diversity of the genomics workforce.
Another experience we’ve had at NIH that I think is very illustrative of this: We recognized that we wanted the African scientists to get more involved in doing genomics. And through a program called H3Africa, the Human Heredity and Health in Africa program, that the NIH and the Wellcome Trust funded, the philosophical mantra is not just to go there and do a bunch of genomics research, but rather empower African scientists to do all the studies and build capacity there. It’s been a success by almost any metric. But it’s exactly what you said: We want them to do the studies, we want them to engage with their local communities. We’ll never build the trust if we just come in and say, “We’re going to do all of this.”
Saey: In terms of privacy, you’ve said a couple of times that you could have somebody’s genome completely sequenced, and then their doctor can use it. Then don’t we get into a situation that could be like the movie Gattaca? Some people could be discriminated against if they don’t have their genetic flaws fixed? Are you somehow creating a class of lesser people and more perfect people who don’t have the genetic flaws that everybody else has?
Green: There are probably about six or seven major ethical dilemmas that you just laid out, and they’re all valid, and we could spend hours talking about each of them. What I would say about our field is we’ve recognized that everything we are doing is a two-edged sword. On the one edge of that sword are these incredible opportunities for improving the practice of medicine. On the other edge of that sword, as with many technologies, it could be used in ways that would be societally unacceptable. It’s a reason why the field has from the beginning always embraced and invested in ethical, legal and social implications research, or ELSI research, which has attempted to anticipate these concerns and try to provide an evidence base to build policies, in some cases, laws.
“Everything we are doing is a two-edged sword. On the one edge of that sword are these incredible opportunities for improving the practice of medicine. On the other edge of that sword, as with many technologies, it could be used in ways that would be societally unacceptable.”
Eric Green
We do have in the United States a major act called the Genetic Information Nondiscrimination Act, which offers some protection against genetic discrimination. We do have laws and policies that protect people’s medical information.
We should recognize that genomics is just part of a bigger set of societal issues, as more and more intimate information about us is electronically available. Trust me, we can learn a lot about you if we just reviewed your Visa card purchases. We as a society have to recognize that, yes, genomic information has some unique attributes, but it’s not totally exceptional. We need to be part of a broader framework for protecting people so that we can benefit from these incredible opportunities.
We just need to make sure we don’t get too far out over our skis. Just because we can do something, doesn’t mean we should. We need to think about all the consequences. We should be constantly understanding what will society tolerate, what do people not want. We also have some things that are going to be completely unacceptable, like doing genetic editing in unborn children. At this stage, we simply don’t think that’s a smart thing to do, we’re not ready to do it, the scientific community has condemned doing it.
Saey: I do want to circle back, because when we were talking about these noncoding sequences, a lot of them help control how genes are used. That may not be so obvious if you just get this string of somebody’s DNA letters. Can you tell from that how those genes will be used? And how those things will be put together? Or is that something you cannot tell by looking at DNA?
Green: There’s no question that sometimes when you talk about genomics, and you talk about genetics, and you focus on the genes — you sometimes see the tree and you lose track of the forest. The forest is medical complexity and biological complexity. And for most things about ourselves, how tall we are, what we look like, and common diseases — hypertension, diabetes, Alzheimer’s, autism, etcetera — things are much more complicated than looking even for one gene. It’s multiple genes. And it’s almost always a greater choreography with our lifestyle, and our social experiences, and our exposures and everything from diet to exercise. There’s a lot more to health and disease than just our genes.
The grand challenge in many ways for the coming decade or two is doing these very large-scale studies where we have as much data as possible, not just genomic data, but lifestyle data and electronic health record data and environmental data and physiological data. There are absolutely going to be patterns. And we’ve just got to find those patterns.
Spotlight
How we got from Gregor Mendel’s pea plants to modern genetics
Philosopher Yafeng Shan explains how today’s understanding of inheritance emerged from a muddle of ideas at the turn of the 20th century.
Saey: We’re almost out of time. It’s been wonderful talking with you. Did we miss anything?
Green: We missed all sorts of wonderful things, but you can only spend so much time walking down memory lane.
What I would say in closing are two things people always need to remember: First of all, how incredibly exciting this field is, and how incredibly eager we are to build our tent with more and more people from all different disciplines. And we also want people of all different populations and ancestral groups from all parts of the world. It’s going to be so important to do that.
The reason we want all these people involved is, we just touched on so many things that we still don’t understand. We need creativity. And we don’t have a playbook. Just like those days where we were bewildered of how we were going to get the Human Genome Project really done, I don’t really know how we’re going to get complete end-to-end understanding of the human genome. But I know if we get creative people working on it, we’ll make incredible progress.
From the archive
-
The Relation of Biology to Physics
With examples related to the cell and chromosomes, Thomas Hunt Morgan calls for the cooperation of physicists in unraveling biological mysteries.
-
Pea Plant Grown by Mendel Given to University
A brief report on the transfer of “one of the most treasured mementos of the modern science of genetics.”
-
All Cancer Tied to DNA
Changes in DNA appear to be “the common denominator of various cancer causes,” Science News Letter reports.
-
Biology Future Predicted
Here’s a summary of pharmacologist, medical historian and prolific writer Chauncey D. Leake’s 10 predictions for the next decade.
-
Inherited Diseases: On the Way Out
“Inherited diseases that plague families for generations will one day be wiped out, scientists now predict.”
-
Gossip, Claret and the Coil of Life
Science News reviews James Watson’s personal account of the discovery of the structure of DNA, The Double Helix, calling it “an important and an unfortunate book.”
-
Molecular Biology’s Flower Child
Joan Arehart-Treichel reports that scientists have deciphered the sequence of DNA subunits that make up a protein-coding gene, and explores what the achievement means for biology.
-
The Gene Idea
On its 60th birthday, Science News looks back at how scientists’ understanding of the units of heredity has changed across the decades.
-
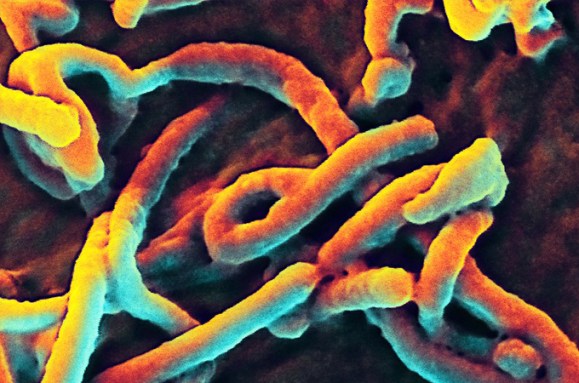
Enter the Virosphere
Reporter Rachel Ehrenberg investigates viruses as masters of manipulation.
-
Factory of Life
Alexandra Witze reports on how synthetic biologists are reinventing nature with parts and circuits.
-
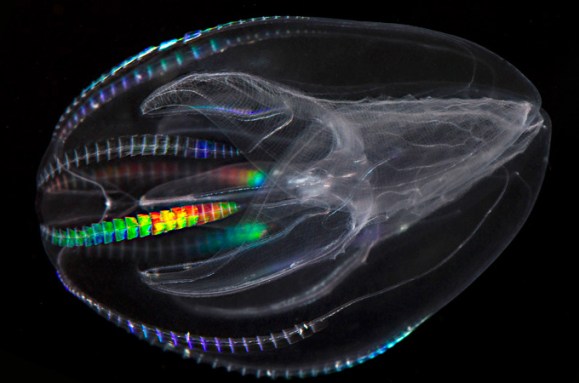
The tree of life gets a makeover
Susan Milius describes how genetic analyses are revamping the tree of life — including putting people and other animals closer to single-celled choanoflagellates than to other multicellular organisms.
-
Special report: Genetic testing goes mainstream
In an award-winning series, Tina Hesman Saey reports on what you can expect to learn (and not) from consumer genetic testing.
The latest
-
We finally have a fully complete human genome
Finding the missing 8 percent of the human genome gives researchers a more powerful tool to better understand human health, disease and evolution.
-
Why experts recommend ditching racial labels in genetic studies
Racial labels don’t explain biological and genetic diversity but do cause stigma. They belong “in the dustbin of history,” a panel of experts says.
-
The new human pangenome could help unveil the biology of everyone
The deciphered DNA includes never-before-explored parts of the genome and better represents the genetic diversity of all humans.
-
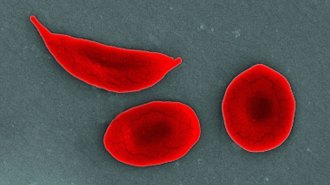
The first CRISPR therapy approved in the U.S. will treat sickle cell disease
In the world’s first CRISPR-based treatment, genetic tweaks to red blood cells aim to help people with the often debilitating disease.