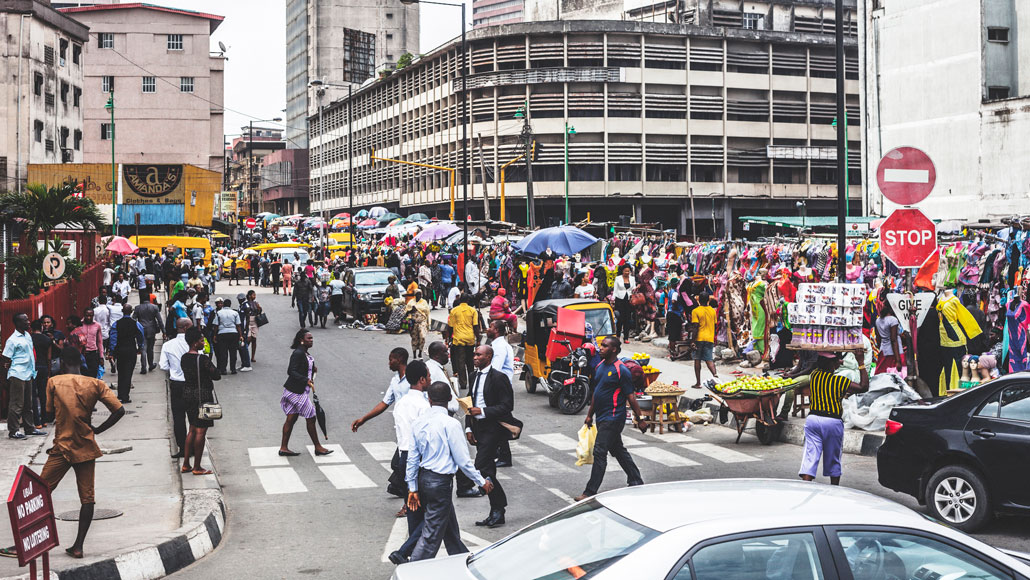
Scientists are trying to build a more complete human reference genome, a master blueprint of our DNA used in biomedical and genetic research. One proposal is to decipher the genomes of 3 million people in Africa (Lagos, Nigeria, shown here).
Peeterv/iStock/Getty Images Plus
As the master blueprint for building humans turns 20, researchers are both celebrating the landmark achievement and looking for ways to bolster its shortcomings.
The Human Genome Project — which built the blueprint, called the human reference genome — has changed the way medical research is conducted, says Ting Wang, a geneticist at Washington University School of Medicine in St. Louis. “It’s highly, highly valuable.”
For instance, before the project, drugs were developed by serendipity, but having the master blueprint led to the development of therapies that could specifically target certain biological processes. As a result, more than 2,000 drugs aimed at specific human genes or proteins have been approved. The reference genome has also made it possible to untangle complicated networks involved in regulating gene activity (SN: 9/5/12) and learn more about how chemical modifications to DNA tweak that activity (SN: 2/18/15). It has also led to the discovery of thousands of genes that don’t make proteins, but instead make many different useful RNAs (SN: 4/7/19). Researcher lay out those accomplishments and others February 10 in Nature.
“That said, the human reference genome we use has certain limitations,” Wang says.
For one thing, it isn’t really finished; gaps remain in the more than 3 billion DNA letter long template, especially in stretches of repetitive DNA. Those are holes where the technology that built the reference doesn’t do a good job of reading every letter. Scientists know there is DNA there, just not how much nor how the letters are arranged. And despite being a compilation of more than 60 people’s DNA, the reference doesn’t fully encapsulate the full range of human genetic diversity.
Sign up for our newsletter
We summarize the week's scientific breakthroughs every Thursday.
Adding diversity
One of the easiest ways to compile a complete catalog of human diversity is to decipher, or sequence, the genomes of 3 million Africans, medical geneticist Ambroise Wonkam of the University of Cape Town in South Africa, proposes in a commentary also published February 10 in Nature. Africa is where modern humans originated, and study after study has uncovered thousands to millions of new genetic variants among people of African descent
For instance, the Human Health and Heredity in Africa project, known as H3Africa, uncovered more than 3 million never-before-seen single letter variants — known as SNPs, short for single nucleotide polymorphisms — by examining DNA of just 426 people from different parts of Africa, researchers reported October 28 in Nature.
Researchers won’t just find single DNA letter, or base, changes when they examine African genomes, Wonkam says. They may discover lots of DNA that no one expected was even in the human genome. Even healthy humans are sometimes missing big chunks of DNA (SN: 10/22/09). And some people may have more DNA than others.
In a 2019 study of 910 people of African descent, researchers discovered an additional 296.5 million DNA bases that aren’t in the current reference. That suggests sequencing Africans might uncover 10 percent or more of the human genome that hasn’t previously been cataloged. That bonus genetic material isn’t necessarily in the gaps researchers already knew about. It hasn’t been found because the 60 or so people whose DNA comprises the reference just didn’t happen to carry it.
“We need a database reference that is representative of humankind,” that is rooted in African origins, Wonkam says. “African population genomic variation is the next frontier” in human genetics.
That doesn’t mean researchers should stop studying people from other parts of the world, he says. A project to examine the genetics of Icelanders, for instance, may uncover genetic variants that arose among the founders of that island nation and are still carried by people today.
But genetic diversity that was present in modern humans before the ancestors of Eurasians left Africa thousands of years ago is still present in people on that continent today, and more variants have arisen as people adapted to specific environments or just by chance.
Research on genetic variation in Africa is sure to help Africans better understand their health problems. But a reference that encompasses the full range of human genetic diversity will help everyone in the world, Wonkam says. Already, new cholesterol-lowering drugs and other medical advances have come from studying the DNA of people of African descent.
Filling in the gaps
While Wonkam’s proposal may solve the genetic diversity problem, it doesn’t necessarily mend gaps in the existing reference genome.
The current reference genome was made by fitting together small strings of DNA like thousands of tiny jigsaw puzzle pieces. In some parts of the genome, the DNA sequence is repeated over and over again, producing virtually identical puzzle pieces. It’s hard to know exactly where all those pieces go and how many repetitions there are. So some repetitive pieces have been left out, leaving holes in the finished puzzle.
That can create problems, Wang says. For instance, doctors may sequence the DNA of a patient and find a genetic variant they suspect might be causing a health problem. But if the suspect DNA isn’t in the current reference, there’s no way to know whether the variant is harmful or not.
“It is time to fully address this problem [with] the limitations of the current human genome assembly,” Wang says. To do that, Wang and other scientists with the Human Pangenome Reference Consortium will use new DNA deciphering technology, called long-range or long-read sequencing, to read each human chromosome from end to end.
In 2020, researchers reported the first fully complete sequence of a human chromosome, the X chromosome. That effort closed 29 gaps in the reference sequence for that chromosome, including 3.1 million bases spanning the centromere, the part of the chromosome important for separating chromosomes during cell division, researchers reported July 14 in Nature. Learning more about centromeres may help researchers understand why chromosome division sometimes goes wrong, leading to cancer or genetic conditions such as Down syndrome.
That early success suggests that long-read sequencing technology can fill in the gaps in the reference genome, and help find the missing 10 percent of DNA. The pangenome team hopes to assemble complete genomes for 350 people from around the world.
And when he says complete, Wang means complete. The reference genome contains more than 3 billion DNA bases, but human cells have more than 6 billion bases. The discrepancy comes from representing just one set of chromosomes instead of the two sets people actually inherit, one from each parent.
That’s because when the DNA was originally sequenced with a person’s DNA being cut into tiny pieces for reassembly later, there was no way to distinguish which little piece came from the chromosome inherited from a person’s mother from the one inherited from the father. So it was all mushed into one.
But by sequencing each chromosome in its entirety, researchers will be able to construct a full picture of a person’s genome, including determining exactly what came from each parent. Those full pictures may allow researchers to better follow patterns of inheritance and track down genetic source of diseases more easily.
Investing in a better reference genome will have big payoffs in other ways too, says Wonkam. The Human Genome Project spent $3.8 billion to build the existing reference. That investment has not only advanced genetic medicine, but has also led to advancements in studying infectious diseases, friendly microbes and other areas of biomedical research.
Having a truly complete reference genome will be even more of a boon, Wonkam predicts. He estimates that the 10-year project to sequence the DNA of 3 million Africans will cost about $450 million a year. But “we’re going to reap a singular benefit, globally, far beyond [the cost].”