Quietly, on the top floor of a nondescript commercial building overlooking Boston Harbor, the future is being born.

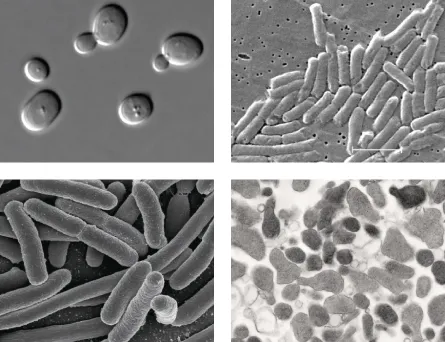

Rows of young scientists tap intently in front of computer monitors, their concentration unbroken even as the occasional plane from Logan Airport buzzes by. State-of-the-art lab equipment hums away in the background. This office, in Boston’s Marine Industrial Park, is what California’s Silicon Valley was four decades ago — the vanguard of an industry that will change your life.
Just as researchers from Stanford provided the brains behind the semiconductor revolution, so are MIT and Harvard fueling the next big transformation. Students and faculty cross the Charles River not to build computer chips, but to re-engineer life itself.
Take Reshma Shetty, one of the young minds at work in the eighth-floor biological production facility. After receiving her doctorate at MIT in 2008, she, like many new graduates, decided she wanted to make her mark on the world. She got together with four colleagues, including her Ph.D. adviser Tom Knight, to establish a company that aims “to make biology easy to engineer.”
Place an order with Ginkgo BioWorks and its researchers will make an organism to do whatever you want. Need to suck carbon dioxide out of the atmosphere? They can engineer the insides of a bacterium to do just that. Want clean, biologically based fuels to replace petroleum taken from the ground? Company scientists will design a microbe to poop those out.
Ginkgo is, in essence, a 21st century factory of life. The researchers working there specialize in synthetic biology, a field that seeks to build living things from the ground up. After envisioning what they want new organisms to do, Ginkgo biologists actually grow vials full of redesigned cells. “We’re going from the place we used to be, in doing science and studying the natural world, to a place where we’re now going to be able to engineer and manipulate it,” says Shetty.
Synthetic biology was born a little more than a decade ago, an offshoot of traditional genetic engineering but distinct in its ambitions, precision and mind-set. Instead of randomly tweaking the genetic blueprints of living organisms and then working backward to identify a cell with a desirable trait, the new field offered the power of designing and building cells with novel functions. Its pioneers dreamed of making armies of organisms that could produce alternative fuels, churn out drugs to battle disease or fill every stomach on the planet by squeezing more food out of each crop acre.
Now, synthetic biologists have laid the groundwork for that radical new future, by building biology’s version of Silicon Valley. One research team has created a new and more complex set of biological building blocks that snap together like Legos, bringing large-scale production of engineered organisms closer to reality. Other scientists have hooked those parts up in a complex living analog of an electrical circuit and programmed it, much like programming a computer. Researchers are now writing code to make cells do things never before thought possible, like hunt down and kill cancer cells.
“This is not just — oh, we’re going to go build something that’s able to make pieces of DNA better,” says Knight, one of the field’s top visionaries. “This is — we’re going to go create a technology infrastructure in the same way that the semiconductor infrastructure was developed.”
From scratch
In its early years, synthetic biology had a less practical, more daring public image. In part that was because of the involvement of J. Craig Venter, the motorcycle-riding, globe-hopping, high-profile iconoclast of modern biology. In the 1990s he led a private effort to decipher the human genetic instruction manual, or genome, that competed with a publicly funded effort. More recently, he sailed his yacht around the world, scooping up water samples every 200 nautical miles to see what microbes were there.
Venter also decided that he wanted to synthesize a living organism from scratch. Such a feat would involve stitching together a creature’s entire genome. DNA’s double helix is made of chains of paired molecules abbreviated as A, T, G and C; long stretches of these letters make up genes, the basic units of heredity. Genes contain the information needed to make proteins, which perform the lion’s share of work in a cell.
Commercial biotechnology companies can easily synthesize short strands of DNA, but putting those together into a full genome is an entirely different matter. So Venter turned to a set of bacteria known as Mycoplasma, which have some of the shortest known genomes (one species has just 580,000 pairs of genetic letters, compared with the 3 billion pairs in the human genome).
Venter’s team took commercially made strands of DNA, then joined them together in his lab using reactive enzymes. After many such steps, the scientists succeeded in fabricating the genome of one Mycoplasma species. The team then inserted the synthetic genome into a second species (which had had its own DNA removed), booting it up. The resulting organism, dubbed “Synthia,” essentially cribbed lab-made DNA to run itself (SN: 6/19/10, p. 5).
Headlines predictably exploded. Life had been made from scratch — sort of. Many synthetic biologists weren’t nearly as excited about Venter’s achievement as the media suggested. These critics point out that his group had simply built an organism to run off a program that already existed in nature; the team didn’t engineer Synthia to do anything new. The crucial difference in today’s synthetic biology, scientists say, is the ability to customize organisms from the start.
“We’re at the beginning of being able to design life in the way that we want,” says Pamela Silver, a biologist at Harvard Medical School and Harvard’s Wyss Institute for Biologically Inspired Engineering.
By design
Engineering new forms of life starts with setting up a biological assembly line, the living equivalent of a transportation innovation. Synthetic biologists aim to reinvent biology in the same way Henry Ford revolutionized automobile manufacturing. Instead of installing standardized spark plugs or carburetors as a car moves down the line, the scientists tuck brand-new biological parts into the body of a bacterium.
To do so, researchers first have to identify distinct, easily defined parts within a cell — biological versions of wheels, hoods, dashboards, engines and so on. Such parts need to be useful in any design, like a power steering pump that works on both a Taurus and a Focus. The parts also need to be standardized so that those made at one factory work with those made at another.
Drew Endy, a synthetic biology pioneer at Stanford, likes to tell the story of William Sellers, who in 1864 argued for the standardization of nuts and bolts so that a wrench made in Wilkes-Barre would fit a nut made in Nashville. Until then mechanics had been working with custom-built hardware. In a lecture at Philadelphia’s Franklin Institute, Sellers called for the country to adopt his new screw design. The standardized, easily measurable shape of its threads would also apply to nuts and bolts, allowing industry to develop a cheap and profitable way to mass manufacture machine shop hardware. Industry agreed, and within just a few years the Sellers screw took off.
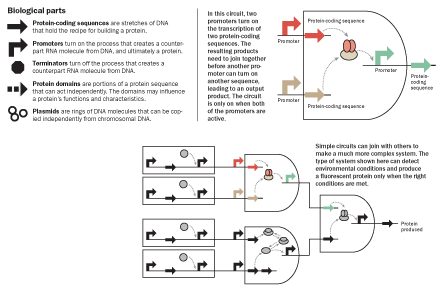
Similarly, scientists are now compiling their own list of biological parts like the Sellers screw. Most parts are stretches of genetic material, much shorter than a gene, that trigger some particular process to turn on or off. A part known as a promoter, for instance, starts the conversion of DNA information into its counterpart, the RNA molecule, while a terminator part stops the action. Many of the parts are proteins known as transcription factors, which hook onto DNA to help control how cells work and respond to their environment.
Scientists make parts by building a stretch of DNA or RNA known to perform a desired job, then adding a standardized string of letters at the beginning and the end to identify it as a part. They then insert the whole thing into a circular strand of DNA until they need it. In 2003, MIT biologists started keeping a formal inventory of these biological parts. Many are added by students who spend summers working on a synthetic biology competition, the International Genetically Engineered Machine contest, or iGEM (SN: 1/12/13, p. 32). Today the list of parts tops 20,000.
Even that roster is too small for some. In his office at Boston University, bioengineer James Collins practically bounces in his chair as he complains about the quality and quantity of most parts. “We just don’t have enough parts to do what we’d like,” he says. “If you survey the original parts out there, we usually use only a dozen or so.”
Collins wants more. Most synthetic transcription factors are designed after versions found in bacteria like Escherichia coli. Collins’ team recently looked instead at yeast cells. Yeast are more complex than bacteria; if engineers could build more parts inspired by yeast, they could use those to create more advanced designs. Working with colleagues including MIT’s Timothy Lu, Collins developed a system to make new transcription factors, and made 19 new ones to start with. “Instead of relying on this small number of things arrived at in nature, we now have a very nice platform that allows you to ramp up and create transcription factors by design, in large numbers,” Collins says. The work appeared last August in Cell.
Cells wired up
Once synthetic biologists have enough parts to work with, the next question is what to do with them. Here, bioengineers take their cue from electrical engineers. Individual biological parts are like the transistors, resistors and capacitors that electrical engineers connect together with wires to create a circuit through which current flows. Circuits can then be connected together on a semiconductor chip to perform computing tasks.
Biologists first reported making synthetic genetic circuits in 2000, when two E. coli papers appeared in the same issue of Nature. In one, a team led by Collins announced the first artificial toggle switch in bacteria; the scientists designed two promoters to interact and drive gene activity if prompted by one molecular signal, and to stop when prompted again.
In the second paper, Stanislas Leibler and Michael Elowitz, then at Princeton, described a synthetic timing switch, in which three genes inhibited one another in sequence, their activity cycling regularly.
These first papers were necessarily clumsy attempts to emulate what nature does effortlessly. But with genetic circuits that accomplished particular tasks, researchers could go one step further: They could connect those circuits with other components, just as electrical engineers do on a computer chip, and program the whole contraption to perform an even more elaborate job.
Across the Charles River from Ginkgo, on the second floor of a gleaming biotechnology building, sits one major hub where biological parts are being turned into sophisticated machinery. This is MIT’s synthetic biology center. Being MIT, it is full of engineers with novel and creative ways to think about programming — even when that programming involves DNA-based circuits rather than electrical ones.
One such tinkerer is Christopher Voigt, whose round face and easygoing manner belie the fact that he commands living organisms to do his bidding. Voigt, a former computer programmer, got into synthetic biology because he saw it as the last frontier. “Being able to write a language that programs E. coli to perform a set of operations is the most challenging problem,” he says.
At first, it wasn’t clear that the dream of programming life would be possible. For most of the 2000s, synthetic biology fought a reputation as being not much more than a bag of parlor tricks. Students working on iGEM teams designed cute proof-of-principle projects, like engineering E. coli to darken in “bacterial photographs” or to smell like wintergreen or banana. It seemed that scientists were connecting and reconnecting biological parts, but not in any kind of profound or truly useful way.
That paradigm is now beginning to shift, Voigt says, as researchers develop more reliable parts and, crucially, many more ways in which to wire them together. Instead of using the same few parts and circuits over and over again, programmers like him now have far more sophisticated designs. “We’re getting to an inflection point,” he says. “Finally.”
In 2012, for instance, Voigt’s research group reported re-creating the main pathway through which bacteria convert atmospheric nitrogen to ammonia. By replacing natural parts with synthetic ones, the scientists essentially adapted the genetic programming guiding the job. The system involved 94 biological parts — a scale of engineering unheard of until recently, Voigt says.
Going one step further to original design, Voigt and his colleagues recently built the largest synthetic genetic circuit to date, described in Nature in November. It involves four sensors, each of which can detect a particular input from the environment. One sensor may detect oxygen levels in a cell, for example, while a second sniffs for glucose. Combining those inputs and others prompts the cell to decide whether to take a particular action.
Voigt and his colleagues hope to use these types of circuits in industrial fermentation vats, so that bacteria inside the vats can sense multiple ways in which the environment changes and adjust activity accordingly. “Some of the very basic circuits are already used in biotechnology, to turn on the production of protein as much as you possibly can,” says Voigt. “But if you’re trying to make materials or chemicals like natural products, that requires a lot more sophistication in terms of timing when things happen.”
Put enough circuits together and program them in the right way, and synthetic biology may soon become a lot more personal. Just as the earliest clunky computers eventually gave way to the iPhone in your back pocket, designer cells might one day become an everyday part of your life. They might even course through your veins — if Ron Weiss has his way.
Weiss works just down the hallway from Voigt at MIT. He began his graduate student career in typical fashion, using computer programs to simulate biological changes in a developing embryo. But then something clicked in his brain. “I remember the day when I thought, let me flip this around,” he says. “Let’s look at what I know in computing and understand how I can program biology.” Then his advisers told him he was too close to getting his Ph.D. to start going down such crazy paths.
Weiss wasn’t going to drop his doctoral quest, but he walked over to Knight’s office and asked to join the budding synthetic biology research group there instead. After many 16-hour days teaching himself how to string together DNA, Weiss changed his focus from engineering to synthetic biology.
Now, in a sort of biological hit job, Weiss’ team has engineered assassin cells to track down and annihilate cancerous cells. The scientists, including Yaakov Benenson formerly of Harvard and now at ETH Zürich, programmed a synthetic circuit that can sense levels of chemicals often found in cancer cells. The circuit also includes a kill switch, a synthetic version of a gene carrying information that can make other cells commit suicide.
Cells carrying this circuit search for cells that are turning cancerous. Once there, the assassin cells flip the kill switch and cause the cancerous ones to off themselves.
In a 2011 Science paper, Weiss’ team showed that this killer circuit could work in human cells in a lab dish. But there’s a long way to go before it could treat cancer in people. Scientists need to find a way to deliver the assassin payload into the body. “We need something like a virus that would go into cells and then compute whether each cell is cancerous or not,” Weiss says. His team is now working to harness a virus that could be used to test the idea in mice. If it works, doctors might eventually be able to inject assassin circuitry into a person suffering from cancer.
Weiss also has his eye on fighting several other important diseases. Diabetes, for instance, can require a person to regularly inject insulin, but Weiss thinks that engineered cells might be able to do that job from within the body. In early theoretical work, his team showed how synthetic gene circuits could steer stem cells to develop into insulin-producing cells. Adding synthetic switches could nudge the insulin production process in one direction or another as needed, the team reported last July in PLOS Computational Biology. The cells could reproduce over and over again, and then die when no longer needed.
Picking up the pace
A medical breakthrough was, in fact, one of synthetic biology’s first major industrial successes: a bioengineered version of artemisinin, a malaria-fighting drug that once had to be laboriously and expensively harvested from the wormwood tree of east Asia. In 2006, researchers from the University of California, Berkeley and Amyris Biotechnologies in Emeryville, Calif., reported that they had engineered baker’s yeast to churn out a crucial precursor to the drug. The scientists teamed up with the pharmaceutical company Sanofi to scale up the process and make the drug in its laboratories. Sanofi is in the early stages of shipping the first commercial artemisinin made using synthetic biology.
Researchers haven’t been as successful with another of synthetic biology’s lofty original goals — to help solve the energy crisis. One early and much-touted promise was that scientists could insert synthetic genes into an organism’s DNA to make it secrete biodiesel or other petroleum alternatives. Some companies, including Ginkgo, are still working on this challenge. But many of the highest profile projects, like those that engineered algae to pump out biofuels, simply haven’t panned out. In most cases, fuel made by synthetically altered organisms can’t compete economically with regular petroleum products.
Most synthetic biologists see this setback as a bump in the road rather than a major derailment for the field. Harvard’s Silver, for instance, has shifted from working on synthetic biology approaches for clean-burning hydrogen fuel to new ways to re-engineer photosynthesis within plants.
Once a molecular biologist, Silver shifted to synthetic biology in the early 2000s so that she could tackle scientific questions no one else could. “The idea of building with biology struck me as very exciting,” she says. Today she oversees one of the largest and most productive synthetic biology research teams, a warren of lab benches and graduate students on Harvard Med’s campus in Boston. Among other efforts, she has developed synthetic genetic counting devices, to keep track of exposures to things like radiation within a cell.
For Silver, synthetic biology is all about accelerating the pace of practical advances. “Biology needs to move faster so that people cheer when something great happens,” she says.
Though it may still lag behind some scientists’ ambitions, there’s no question the field is progressing rapidly. Time and again, researchers have invented new methods for assembling synthetic parts and genetic circuits cheaper, faster and more easily than before.
In 2009, scientists working for Venter came up with a new way of stitching together different biological parts by using DNA strands with overlapping letter sequences on their ends. Biologists can easily add the matching sequences to any parts they want to link, then stir in some enzymes and, voilà, assembly. The method, invented by Daniel Gibson, has caught on quickly because it lets scientists patch together more than a dozen DNA strands at once. Just a year after its invention, Gibson assembly inspired a devotional YouTube video from an iGEM student team. Today it is used in nearly every synthetic biology lab.
And at Harvard, biologist George Church devised a technique that makes multiple changes to an organism’s genome at a time. MAGE (for multiplex automated genomic engineering) is like a genetics editor on speed; it zips through, finding and tweaking DNA automatically so that researchers can add various synthetic components at once and test what they do. In 2011, Church and colleagues founded a company, Warp Drive Bio in Cambridge, to use a version of this superfast technique to hunt for potential new drugs in natural compounds.
The market for synthetic biology products is still quite small, and one of Church’s earlier start-ups failed after trying to do too much too fast. But he and other visionaries are convinced that synthetic biology will be big. Huge, in fact — as huge as the Internet.
And they should know. Several scientists pushing the field forward today are former electrical engineers who helped develop key components of what became the Internet, such as its ARPANET predecessor.
“The Internet disrupted the world — it was unleashing a completely different aspect of nature,” says MIT’s Randy Rettberg, a former engineer at Sun Microsystems who now runs the iGEM competition. So, too, will synthetic biology, by telling biological matter precisely how to behave. “First we had the industrial revolution, then we had the network revolution,” says Rettberg, “and now we have the matter revolution.”
Rettberg thinks that synthetic biology’s full impact, like that of the Internet, will take decades to emerge. “We’re only about 10 years into it; it took about 25 years from ARPANET until you had the beginning of the World Wide Web,” he says. “And although the Internet took a very long time, its impact was dramatically bigger than everybody but the visionaries imagined.”
It’s hard not to get caught up in Rettberg’s enthusiasm as he bustles about the iGEM offices in Cambridge, proudly introducing students who help box up test tubes full of biological parts and mail them out to competitors. This is a man who charted out the final phase of his scientific career on graph paper to see if he had enough time left to learn something completely new. Then he taught himself synthetic biology.
As did Rettberg’s longtime friend Knight, also a former electrical engineer. Knight now spends most of his time at Ginkgo’s offices, where his business card reads simply “DNA Hacker.” As automated machines whir in lab space across the hall, testing what freshly engineered organisms can do, Knight dreams up new designs for Ginkgo to try. Just as he once dreamed up what would become some of the first single-user computer workstations.
“I knew this was the exciting thing to go do,” he says. “What does it take to make the next Intel? I am actually interested in making that work.”
Solving hunger
Synthetic biology may help farmers feed more people. For millennia, crops have been bred with an eye toward improved harvests. Later, genetic manipulations upped plant yields and made crops more resilient against drought and other hazards. Now, scientists are looking at tweaking photosynthesis. “You don’t need to increase the biomass of plants by that much to solve the food problems across the world,” says Harvard’s Pamela Silver. One idea is that new enzymes could boost the amount of energy that plants can extract from the sun. Another suggests there might be a totally different way to pull usable carbon from the atmosphere. In the April 2012 Applied and Environmental Microbiology, Silver and colleagues reported engineering a bacterium to churn out up to 200 percent of its initial cellular mass as sugar. The work could be used to develop plants that produce more food per harvest.
Making energy
An early hope for synthetic biology was that it could wean society off fossil fuels. Engineering microbes to churn out hydrocarbons would presumably be a lot cleaner and more climate-friendly than extracting and burning coal and oil. Since 2000, the U.S. Department of Energy has poured millions of dollars into funding synthetic biology biofuels research, such as new types of algae to secrete biodiesel or other engineered fuels that don’t have to be pumped from the ground. So far, progress has been limited.
Treating patients
One of the most obvious goals of synthetic biology is to make people healthier. Engineering new drugs, or designing cells that can target disease inside the body, has been a goal of the field from the start. An early success involved creating a bioengineered version of a drug to fight malaria. Researchers managed to engineer a species of yeast to produce large amounts of a chemical precursor to the antimalarial drug artemisinin, typically harvested from the wormwood tree of east Asia. The pharmaceutical company Sanofi is now working to bring the process to market. In another take on better health, engineered human cells could locate and eliminate cancerous cells by tricking the evildoers into committing suicide. Though the technique has been demonstrated in a lab dish, it is still far from tackling cancer in real human patients.
Cleaning up
Microbes are already used at oil spill sites, eating petroleum components and converting them into less hazardous by-products. Designing synthetic versions that can do the job quicker, and perhaps break down more stubborn pollutants such as pesticides and radioactive waste, would be a logical next step. Researchers at Spain’s National Center for Biotechnology have designed circuits capable of redirecting microbes to feast on industrial chemicals instead of sugar.
Reviews give green light, encourage caution
Engineering life is not the sort of thing you can do quietly.
Ever since biologists first started piecing together genetic components, ethicists have pondered the implications. Could an artificial form of life turn out to have unexpected consequences, like invading the environment or otherwise running amok? And what about bioterrorists who might want to get their hands on synthetic bugs and put them to nefarious uses?
A March 2012 report from Friends of the Earth, the International Center for Technology Assessment, and the ETC Group — nongovernment organizations that have worked against genetically modified organisms, among other causes — calls synthetic biology “an extreme form of genetic engineering” that is “developing rapidly with little oversight or regulation despite carrying vast uncertainty.” Not since the 1990s’ birth of nanotechnology, the engineering of the very small, has a new technology elicited such ire.
Nearly every major safety review of synthetic biology, though, has given the field a cautious green light. A 2010 government review, requested by President Obama after Craig Venter booted up a cell with a synthetic genome, suggested there was no need to create a new government body to oversee synthetic biology research. Rather, the report’s authors promoted the idea of “prudent vigilance” — paying attention to what’s happening in the field without regulating it out of existence from the start. “With these unprecedented achievements comes an obligation to consider carefully both the promise and potential perils that they could realize,” the report said.
The Woodrow Wilson International Center for Scholars in Washington, D.C., has also started a scorecard for tracking public discussions about synthetic biology. An update last July found that many U.S. federal agencies had begun taking steps to learn more about the field, as recommended by the presidential report. Still, the center says, more work is needed.