PROTEIN POWER Proteins from Staphylococcus aureus bacteria make amyloid-like clumps (the streamerlike structures in this illustration) that are toxic to human cells.
Dima Abelski/Dimedia
Clusters of a toxic bacterial protein have a surprising structure, differing from similar clumps associated with Alzheimer’s and Parkinson’s in humans, scientists report in the Feb. 24 Science.
These clusters, called amyloids, are defined in part by their structure: straight regions of protein chains called beta strands, folded accordion-style into flat beta sheets, which then stack up to form a fiber. That definition might now need to be broadened.
“All the amyloids that have been structurally looked at so far have certain characteristics,” says Matthew Chapman, a biologist at the University of Michigan in Ann Arbor who wasn’t part of the work. “This is the odd amyloid out right now.”
In the human brain, misfolded proteins can form amyloids that trigger neurodegenerative diseases. But amyloids aren’t always a sign of something gone wrong — some bacteria make amyloids to help defend their turf.
In Staphylococcus aureus, for example, the PSMα3 protein assembles into amyloids that help the bacteria kill other cells. Previous research suggested that PSMα3 clusters were like any other amyloid. But researchers using X-ray crystallography found that instead of straight beta strands, the PSMα3 fiber was made up of curly structures called alpha helices that resemble an old-fashioned phone cord. The helices still formed a familiar fiber shape just like the beta strands did, but the sheets making up that fiber were rippled instead of flat.
Story continues below image
Atypical amyloid
A protein from Staphylococcus aureus bacteria makes amyloids from sheets of curly alpha helices (left) rather than the typical flat beta strands (right).
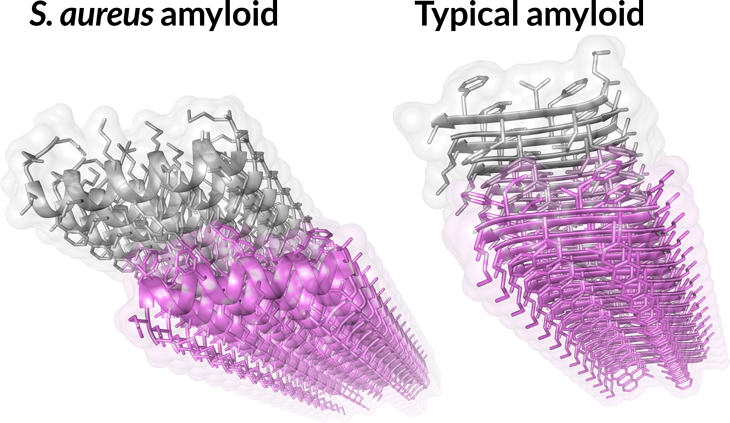

“From the outside, it looks exactly the same. But zooming in, it looks completely different in its fundamental units,” says study coauthor Meytal Landau, a biologist at the Technion–Israel Institute of Technology in Haifa. “It’s something that really threw me off.”
Chapman first thought the finding must have been a mistake. Alpha helices are a very common protein structure, but “they don’t typically stack on each other to form a sheet,” he says. “That’s a really rare structure in nature.” Now, he says, it’s quite clear that alpha helices are building this amyloid.
Getting such a detailed picture of an amyloid is a challenge, so Landau isn’t sure whether PSMα3 is truly an exception or whether other proteins can make similar amyloid-like clusters.
Either way, knowing the structure can be useful. When Landau and her colleagues prevented the PSMα3 from forming an amyloid-like structure, the bacteria weren’t as good at attacking human immune cells. Rather than hunting for strong drugs that target the hard-to-kill bacteria themselves, Landau suggests, scientists might eventually be able to develop drugs that target the proteins controlling the bacteria’s aggression.






