In their everyday battles against harmful bacteria, physicians, food producers, and others need to know quickly which foe they’re facing. Yet the procedures currently used to identify bacterial colonies are often time-consuming and expensive.
Technicians must wait hours or days for suspect bacteria to grow into a colony for testing in a laboratory. However, by taking a picture of a colony with laser light, a new technique created by Bartlomiej Rajwa of Purdue University in West Lafayette, Ind., and his colleagues identifies the colony without further delay.
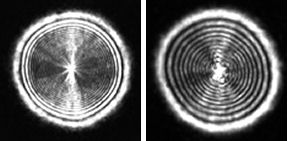
In their method, the researchers shine a laser beam through a petri dish dotted with bacterial colonies, project an image of one colony at a time on a screen, and record that image with a digital camera. This approach requires no stains or costly custom chemicals, the team reports. The components of the prototype device are inexpensive, so the technology could be widely affordable.
The researchers describe their prototype system and initial results in the current, May/June Journal of Biomedical Optics.
Identifying tiny biological entities from patterns of scattered light isn’t new. For instance, widely used machines called flow cytometers distinguish and count various human-cell types by using light scattered as cells pass through a laser beam, notes biophysicist Robert M. Zucker of the Environmental Protection Agency in Research Triangle Park, N.C.
To differentiate colonies of bacteria about 2 millimeters across by the complex light patterns they create, the Purdue team devised a method similar to automated face recognition. The researchers recorded the pattern of light emerging from each bacterial colony and compared it with 120 mathematically derived shapes. In that way, they generated a classification code for each colony.
Similar codes identified colonies made up of the same species of microorganisms, Rajwa explains. The researchers don’t hypothesize which structural features of bacteria or their colonies cause the differences in the patterns.
In tests described in the new report, the system identified with about 70 percent accuracy worrisome colonies of Listeria—a genus of foodborne bacteria in which only some species are harmful.
Zucker calls the new technique “promising” but not yet accurate enough.
Al Brunsting, an engineer with Panduit Corp. in Orland Park, Ill., says that he’s “skeptical” of the approach. One concern is that its developers haven’t shown that a colony’s pattern would be unchanged if measured under varying conditions.
Rajwa says that the pattern-recognition algorithm accommodates considerable variability. What’s more, he adds, his team has recently upgraded its system to a version that’s “vastly superior” to the one reported in the journal. The new system classifies Salmonella, Vibrio, Bacillus, and other bacteria. It achieves 98 percent accuracy for many types of bacteria, Rajwa says.






