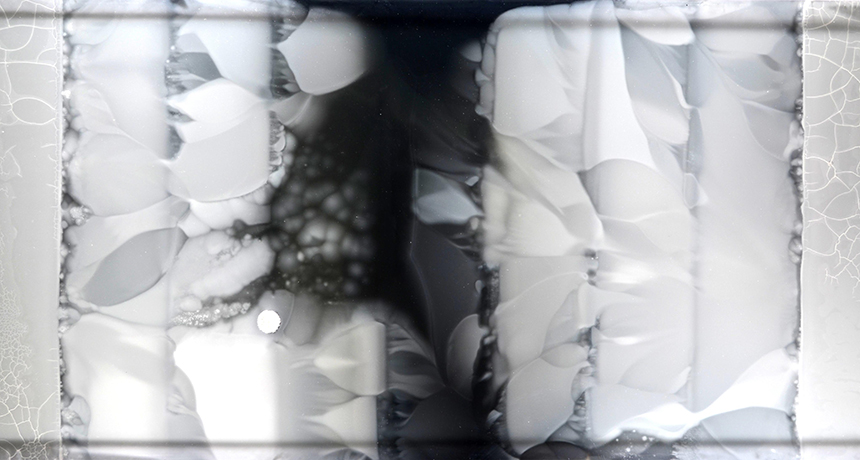
PETRI PLATTER A petri dish more than a meter long helped scientists visualize the evolution of antibiotic resistance in E. coli bacteria. Bacteria placed on the outer edges had to adapt to higher and higher levels of antibiotics as they moved toward the center of the plate.
Harvard Medical School