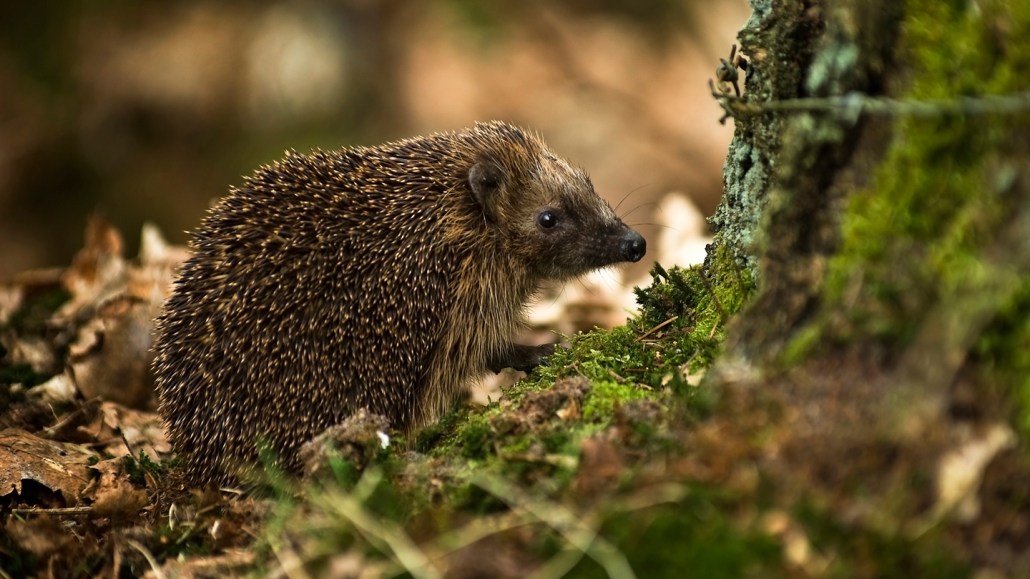
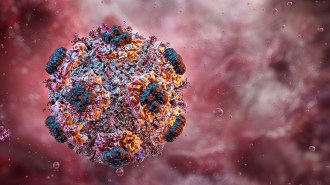
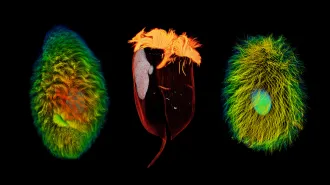
Workers at wildlife rescue centers swabbed the noses, skin and feet of hundreds of European hedgehogs (Erinaceus europaeus shown), revealing that many of the critters harbored a type of MRSA superbug.
Hrald/Wikimedia Commons (CC BY-SA 3.0)
Beneath the prickly spines of European hedgehogs, a microbial standoff may have bred a dangerous drug-resistant pathogen long before the era of antibiotic use in humans.
It’s no question that antibiotic use accelerates drug-resistance in bacteria that colonize humans, says Jesper Larsen, a veterinarian at Statens Serum Institut in Copenhagen. But, he says, these microbes had to get the genes to give them resistance from somewhere, and scientists don’t know where most of these genes come from.
Now, for one type of methicillin-resistant Staphylococcus aureus, or MRSA, Larsen and colleagues have tracked its evolution to hedgehogs hundreds of years ago. On the skin of these critters, a fungus that produces natural antibiotics may have created the environment for drug resistance to evolve in the bacteria, the researchers report January 5 in Nature.
One of the most common drug-resistant pathogens, MRSA infects hundreds of thousands of people worldwide each year, and these infections can be hard to treat. The specific type of MRSA that the new study focuses on causes a fraction of the cases in humans.
The team first found MRSA in hedgehogs by coincidence years ago when biologist Sophie Rasmussen, who was part of the new work and is now at the University of Oxford, approached Larsen’s team about sampling a freezer full of dead hedgehogs. Of these animals collected from Denmark, 61 percent carried MRSA. “We found this extremely high prevalence in hedgehogs,” Larsen says, suggesting that the animals were a reservoir for the drug-resistant superbug.
In the new work, the scientists surveyed hedgehogs (Erinaceus europaeus and Erinaceus roumanicus) from 10 European countries and New Zealand. Workers at wildlife rescue centers swabbed the noses, skin and feet of 276 animals. MRSA was prevalent in hedgehogs in the United Kingdom, Scandinavia and the Czech Republic.
Analyzing the S. aureus, the team found 16 strains of mecC-MRSA, named after the gene that confers resistance, and mapped the evolutionary relationships between them by comparing mutations across their genetic instruction manuals, or genomes. From the analysis, the team inferred that the three oldest lineages emerged 130 to 200 years ago in hedgehog populations, periodically infecting people and cattle long before penicillin hit the market in the 1940s. Hedgehogs may be the source of nine out of the 16 lineages, the researchers report.
“There is no doubt that our usage of antibiotics is the main driver of resistance in human pathogens,” says Anders Larsen, a microbiologist at Statens Serum Institut who was also was part of the team. “This is a very special case where we can just track it back to an origin.”
But that doesn’t explain how the hedgehogs’ S. aureus developed resistance. The team got a clue from a 1960s research study about Trichophyton erinacei, a fungus that causes “hedgehog ringworm” in humans. That study reported that T. erinacei on hedgehog skin killed some S. aureus but not others that were resistant to penicillin. Growing T. erinacei in the lab, the researchers identified two penicillin-like antibiotics pumped out by the fungi.
This finding suggests that hedgehogs are a MRSA reservoir because “they’re living cheek by jowl with organisms that are producing penicillin,” says Gerry Wright, a biochemist at McMaster University in Hamilton, Canada, who was not involved with the study.
The fungi “live in a bad neighborhood,” Wright says. They have to compete with other microbes, such as S. aureus, for resources and a spot to colonize on the host, and “they have to work out this arrangement where they can protect themselves.”
You can’t think about antibiotic resistance without considering environmental connections, Wright says. The evolution of resistance is a gradual process shaped by natural selection, he says. Wright’s work has shown that in places that have escaped human influence, antibiotic resistance has ancient origins. People have searched for this evolution mostly in the soil microbial community, or microbiome (SN: 2/14/06). But the microbiomes of animals provide another potential source for the genes that confer resistance as well as for sources of new antibiotics, he says.
The history of antibiotics in the last century is a cycle of new drug discoveries shortly followed by microbial resistance cropping up to those drugs. That shouldn’t be a surprise, Wright says. “Because antibiotics have been on the planet for billions of years, and resistance is billions of years old,” he says. If scientists don’t better understand where resistance comes from, even as researchers discover new drugs, he says, all we’ll be doing is playing catch-up.