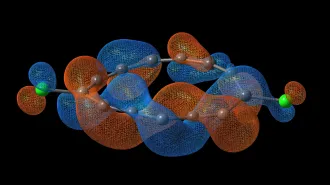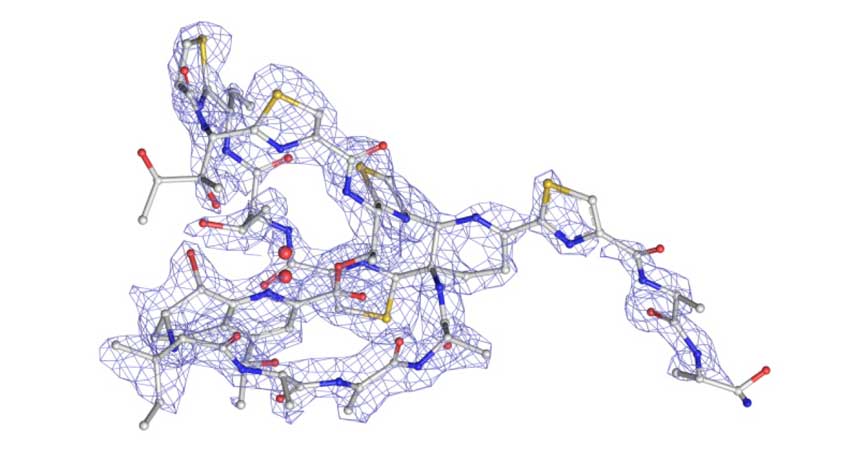
SPITTING IMAGE In just a day, researchers using a new technique to identify a substance’s chemical structure had mapped out the antibiotic thiostrepton (carbon atoms are gray; nitrogen, blue; oxygen, red; and sulfur, yellow). With current methods, that mapping could have taken months or years.
C.G. Jones et al/ChemRxiv.org 2018
The one-hour photo booth has met its molecular match.
By adapting a technique for determining protein structures, two independent teams have charted chemical structures of antibiotics, hormones and other compounds with unprecedented speed. Depending on the molecule, it took between 30 minutes and a day to determine structures, where traditional techniques could take months to years.
The approach could help forensic chemists and sports officials identify illicit drugs and accelerate drugmakers’ pursuit of new ones. The work was published online October 16 in Angewandte Chemie and October 17 at ChemRxiv.org.
“I haven’t been this excited about a finding in chemistry in a long time,” says organic and medicinal chemist Donna Huryn of the University of Pittsburgh who was not part of either study. “It’s going to change the way everybody works.”
For molecular scientists, it’s essential to know the atom-by-atom connectivity of what they’re working with. It’s how pharmaceutical companies verify drugs’ composition and purity, for instance. But most techniques for discerning chemical structures are indirect. It’s like having a fingerprint, an artist’s sketch and dental records when what you want is a photo, explains Brian Stoltz, an organic chemist at Caltech and a coauthor of the ChemRxiv study.
The closest thing to a camera for molecules is a technique called X-ray crystallography. It involves bombarding a crystal — tiny rock candy made from many copies of one molecule stacked together — with X-rays, and calculating a structure based on how the X-rays bounce off the atoms making up the crystal. But creating big enough crystals for this approach, at least 50 micrometers on one side, can take years and doesn’t always work.
Powder to picture
A sample from a decades-old bottle of progesterone (left), applied to a Swiss cheese–like sample holder (middle left), produces a pattern after electron bombardment (middle right) in less than three minutes that can be translated into its chemical structure (right). The whole process for progesterone took about 30 minutes.
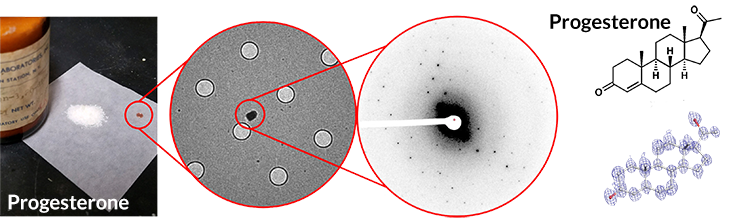
To avoid that size bottleneck, the two teams fired electrons at molecules instead. X-rays mostly pass through matter, whereas electrons bounce off matter more readily, says Tamir Gonen, an electron crystallography expert at UCLA and a coauthor of the ChemRxiv study. So it’s possible to get structures from roughly 100-nanometer crystals, far smaller than those needed for X-ray crystallography. Such crystals are invisible to the naked eye, but readily available in bottles of chemicals straight off the shelf.
In the Angewandte study, Tim Gruene of the Paul Scherrer Institute in Villigen, Switzerland, and his team cracked open a capsule of cold medication containing a mixture of active and inactive ingredients. And, from a speck of powder, the researchers determined the structure of acetaminophen inside. That previously solved structure isn’t exciting, Gruene says, but the fact that it came from a mishmash of ingredients is. While not every molecule will produce the tiny crystals needed, crime labs analyzing trace samples could find this technique handy. It could also be useful in contested drug patent cases, because the method unambiguously verifies structures from crushed-up pills, Gruene says.
Using electrons to map structures isn’t a new idea. The 2017 Nobel Prize in chemistry went to scientists who used this concept to map proteins with hundreds of thousands of atoms (SN: 10/28/17, p. 6). And a few researchers previously obtained structures of organic molecules with a few dozen atoms, but those results didn’t catch on among chemists.
By making the value of this technique to chemists clear, these studies have provoked wonderment on chemistry blogs and social media. It took a couple of years for the team to overcome technical hurdles, says UCLA organic chemist Hosea Nelson, but once it had, it obtained six or seven structures in one afternoon.
At that point, Nelson recalls, “we said — with expletives mixed in — that this was the most exciting day as scientists that we’d ever experienced.”