Opiates for the masses may not be far off. Scientists have figured out two of the final steps in the chain of chemical reactions that synthesize morphine in the opium poppy.


Pinpointing the cellular workhorses and the genes involved in making morphine may lead to new production methods for the drug and its chemical cousins such as codeine, oxycodone and buprenorphine, scientists report in a paper published online March 14 in Nature Chemical Biology.
Morphine and its relatives, widely used as painkillers in developed countries, are fairly expensive and are often taken for extended periods of time. The new research may lead to better ways of engineering yeast or other microbes to make these painkillers — perhaps skirting the social and political morass of agricultural poppy production, the source of heroin.
“Moving production of morphine and its metabolites such as codeine into a microbial system — if you could get yields up — could help lower costs,” says bioengineer Christina Smolke of Stanford University, who was not involved in the research. Instead of having to purchase these opiates from other nations, “maybe countries could even do local synthesis,” she says.
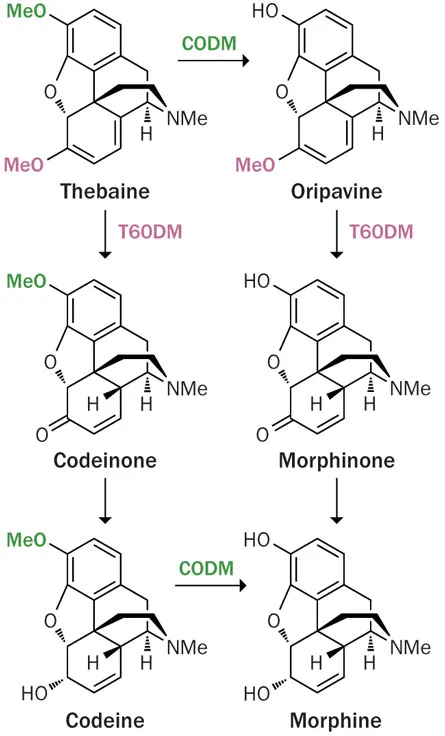
The new work identifies two enzymes — the proteins that cells use to build molecules and make reactions go — involved in turning the chemical precursors thebaine and codeine into morphine. Study coauthors Jillian Hagel and Peter Facchini of the University of Calgary in Canada also pinpointed the genes encoding each enzyme and verified this genetic role with poppy plant experiments.
“This is really terrific work,” says Philip Larkin, head of the plant product metabolic engineering program at Australia’s national science agency CSIRO in Canberra. “Having these genes in the hand gives you much greater versatility.” For example, scientists could engineer high-yield plants by cranking up the activity of the morphine synthesis genes, Larkin says.
Scientists could also block morphine production with engineered viruses that shut down the genes. In theory, such viruses might be used to eradicate opium poppy crops in places such as Afghanistan. But narcotic control experts question the wisdom of such a maneuver.
“There are formidable tactical obstacles that would have to be addressed,” says Charles S. Helling, former senior scientific advisor to the State Department’s Bureau of International Narcotics and Law Enforcement Affairs. “But the even bigger problems are political,” he adds. “It’s a very difficult situation that is further complicated by the military situation.”
Morphine is an alkaloid, a class of compounds characterized by a ringed molecular structure incorporating a bit of nitrogen. “Among all the natural products, alkaloids tend to display the most potent pharmacological effects,” Facchini says. Plants produce roughly 12,000 kinds of alkaloids, including nicotine, strychnine, caffeine, mescaline, quinine and atropine.
A handful of very old plant groups, including the poppy and buttercup families, produce the class of alkaloids that morphine belongs to, called benzylisoquinoline alkaloids. The main building block for the roughly 2,500 alkaloids in this class is the amino acid tyrosine. A 15- to 20-step reaction pathway turns tyrosine into morphine. While questions remain about some of the very early reactions, pinning down the final morphine production steps is the key to unlocking a host of practical applications.
Years of research, gift plants, a bit of luck and the “Herculean effort” of then graduate student Hagel led to the discovery, says Facchini.
The researchers began with three high-morphine varieties of opium poppy, Papaver somniferum, and a mutant plant that makes the morphine precursors thebaine and oripavine but can’t make morphine itself. Hagel constructed an enormous DNA library from these plants, which the team used to determine which genes were turned on in the morphine-making poppies. She then compared this activity to that of the mutant plant that couldn’t put morphine together.
After determining the genetic blueprints of the genes that differed, Hagel and Facchini checked those DNA sequences against a database to reveal the enzymes’ identities. To verify the enzymes’ role in making morphine, Hagel stuck one of the genes into the bacterium E.coli, put the critter in a flask with some thebaine, and left it overnight.
“When she came back the next morning, the thebaine was all gone,” says Facchini. “That’s when her eyes got big…. Finding it all had been turned into morphine — that gives a grad student a great sense of power, when they can make morphine.” The scientists dubbed the enzymes thebaine 6-O-demethylase and codeine O-demethylase.
Both of the newly identified enzymes are in charge of the same structural task — removing a methyl group, a common chemical ornament comprising a carbon and three hydrogen atoms. But in the hunt for these morphine-synthesis enzymes, many scientists were led astray. There was an assumption that poppies used a methyl-removing enzyme similar to the one that the human liver uses to remove methyl groups. But poppies use enzymes from an entirely different class, the researchers report.
“These are enzymes that have eluded discovery for a long time,” says MIT biochemist Sarah O’Connor. And they turned out to be enzymes that weren’t really on the radar. “In plants, it’s very hard to figure out the enzymatic steps of a pathway,” she notes. “This is a beautiful example of how you can use modern molecular biology tools to solve this problem.”







