Cue the corny jokes.
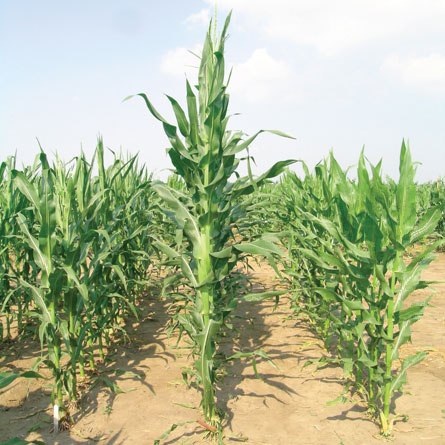


Researchers have completed a draft of the maize genome. And while people may quip about the “amaizeing” achievement, scientists say the genetic blueprint of one strain of corn reveals serious amounts of genetic diversity and some weighty biology lessons that could lead to improvements in the already economically important crop plant.
A series of research papers published in the Nov. 20 Science and in the online journal PLoS Genetics report the genome draft and analyses of the plant’s genetic makeup. The work, conducted by many institutions and funded by the National Science Foundation, reveals that corn has an unusual ability to make new genes, lose others, alter activity of its genes and withstand radical genome remodeling. And surprising differences between two strains of corn may provide clues about why hybrids sometimes grow or yield better than parent strains.
Scientists have known that different strains of maize can vary widely in their genetic makeup. Although corn was domesticated only 8,000 to 10,000 years ago from the grass teosinte, the genetic diversity between any two strains of corn exceeds that found between humans and chimpanzees, species separated by millions of years of evolution. For instance, the strain B73, the agriculturally important and commonly studied variety decoded by the maize genome project, contains 2.3 billion bases, the chemical units that make up DNA. But the genome of a strain of popcorn decoded by researchers in Mexico is 22 percent smaller than B73’s genome.
“You could fit a whole rice genome in the difference between those two strains of corn,” says Virginia Walbot, a molecular biologist at Stanford University.
Much of that difference is due to the action of transposable elements, better known as jumping genes. Transposable elements (transposons for short) are mobile pieces of genetic material that hop around the genome, sometimes taking genes or pieces of genes with them. It was in corn that Nobel laureate Barbara McClintock discovered transposons. The genome project discovered some new families of transposons, revealing a total of 1,300 such families in maize, says Patrick Schnable, a maize geneticist at Iowa State University in Ames. Together, transposons and other repeated pieces of DNA make up 85 percent of the maize genome in the B73 strain.
That’s an unusually high volume of repetitive DNA, and it made the decoding effort challenging, says Richard K. Wilson of the Genome Center at Washington University School of Medicine in St. Louis. “This was a really tough one.” The large number of repeated stretches meant the popular shotgun approach — relying on computers to assemble a scatter-shot of DNA fragments from all over the genome into a coherent picture — wouldn’t work for deciphering corn’s code.
Instead, researchers had to fall back on a slower, bit-by-bit approach similar to that used in the Human Genome Project, first putting the pages of corn’s genetic instruction book in order and then determining the DNA letters on each page.
The project cost $30 million and is not entirely finished, but is already providing maize researchers with a wealth of knowledge.
B73’s genome contains about 32,000 genes, the researchers report in Science. Transposons have also left bits and pieces of genes scattered throughout the genome. Some of those fragments are active and may help control activity of other genes, Schnable says. The researchers also uncovered evidence that maize strains are creating new genes and losing others. For example, at least 180 of B73’s genes are missing from another strain of corn known as Missouri 17 or Mo17. In fact, thousands of pieces of DNA found in one strain are completely missing from the other, Schnable and colleagues report in one of the 10 companion articles published in PLoS Genetics. This degree of structural variation is not seen in any other plant, animal or fungus studied so far, says Schnable.
The researchers also describe variation in the number of copies of genes the two strains carry. Some of those gains and losses of genes and other sequences might contribute to hybrid vigor, a condition in which offspring are heartier and better yielding than either parent.
Researchers don’t yet know the source of hybrid vigor, but the genome sequence may make it easier to trace, Schnable says. To determine whether hybrid vigor is a product of gene activity, he and others compared gene activity in offspring of crosses between the B73 and Mo17 maize strains. In thousands of the hybrid’s genes, only the copy inherited from the male parent was active, the team reports in Science. This genetic phenomenon of only one parent’s gene being active is called imprinting. In humans, less than 1 percent of genes are imprinted. The magnitude of the imprinting in maize — the number of genes and the strong tendency for only paternal genes to be active — is surprising, Schnable says.
The researchers don’t know how or if imprinting contributes to hybrid vigor.
Corn’s liberal genetic policies might eventually get it into trouble. For instance, “In other species, it’s a rare individual that has active transposons,” Walbot says, because active transposons have the potential to disrupt crucial genes. But the strategy has worked for corn, allowing its genome to nimbly adapt to changing environmental conditions, often from generation to generation. But, the species could also, eventually, split into multiple species.
“Corn is living in peril, you might say,” Walbot says. “It has these features that allow flexibility in its genome, but there’s a cost to running this game.”