Genes alone don’t make the man — after all, humans and chimps share roughly 98 percent of their DNA. But where, when and how much genes are turned on may be essential in setting people apart from other primates.
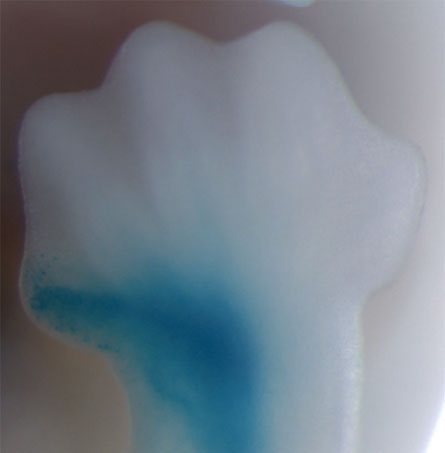
A stretch of human DNA inserted into mice embryos revs the activity of genes in the developing thumb, toe, forelimb and hind limb. But the chimp and rhesus macaque version of this same stretch of DNA spurs only faint activity in the developing limbs, reports a new study in the Sept. 5 Science.
The research supports the notion that changes in the regulation of genes— rather than changes in the genes themselves — were crucial evolutionary steps in the human ability to use fire, invent wheels and ponder existential questions, like what distinguishes people from our primate cousins.
“We’re trying to find out what makes us human,” says geneticist James Noonan of YaleUniversity, who led the study with colleagues from Lawrence Berkeley National Laboratory, the Genome Institute of Singapore and the Medical Research Council in the United Kingdom. “We know that the things that make us human biologically are encoded in there somewhere.”
Noonan and colleagues combed through the vast regions of human DNA that do not contain code for making proteins. Formerly dissed as “junk DNA,” sections of these non-gene regions are now known to play a regulatory role, dialing down or cranking up the activity of actual genes.
Like electrical wiring in a house, genes may be turned on in many places at once, even though they might only be needed in one area, such as the eye, comments Francesca Mariani of the Broad Center for Regenerative Medicine and Stem Cell Research at the University of Southern California,. So while the new study can’t say what these regulatory changes might do in a human embryo, “this does show how a few small changes could make a big difference” she says. “The thing that is intriguing about this is the direction it could go.”
The researchers identified a DNA region made up of 546 base pairs, or “letters” of code. While this region of DNA has barely changed during the evolution of backboned creatures, it has accumulated 16 changes since the ancestors of chimps and humans split, some 6 million years ago, the researcher report. Thirteen of these changes were clustered within an 81 base-pair region.
“If you slip into our shoes as genetic detectives, that many changes in a small area calls out for investigation,” says Sean Carroll of the University of Wisconsin–Madison.
To see the effects of these changes, the researchers inserted the human version, the chimp version, and the rhesus macaque version into mouse embryos. All versions turned on genes in the developing ear and eye, and in the embryonic gills known as pharyngeal arches.
But only the human version boosted the activity of genes in the mouse’s primordial thumb, forelimb, hind limb and toe, the researchers report.
Such changes in gene activity could have led the developments that allow humans to walk upright, such as shorter toes, the researchers speculate. But much more work needs to be done to establish such a link, cautions Carroll.
“We can’t put full flesh on its meaning just yet. One has to expect that in the remodeling of human limbs, brain or skeleton that many, many, many genes were involved over a long period of time,” says Carroll. “If you consider this a piece of the genetic puzzle, we don’t know how big this piece is, or where it fits in.”
Nonetheless, he calls the new work “pretty enticing.” Scientists trying to get at the genetic and developmental changes that make humans human are building on decades of work with model organisms such as fruit flies and mice, where swapping genes around to understand their activity is old hat. Many of these experiments just can’t be ethically done on humans, Carroll notes.
Noonan and his team speculate that the changes in the human version of this region of code might make it easier for certain proteins known as transcription factors to attach to the DNA. Transcription factors kick these regulatory stretches of code into gear.
Carroll notes that there are other ways to spur the regulatory stretches into action. Work by Carroll published in Cell in August found that the “letters” of code aren’t all that determines how aggressively transcription factors attach to the regulatory regions. The actual spacing of the docking regions where the proteins attach can also influence the activity of the regulatory regions.
“There are many ways to tinker with them and get a different output,” he says.