Charles Darwin didn’t know about genes and DNA. In fact, hardly anyone noticed when Gregor Mendel, a monk whose pea experiments eventually led to modern genetics, published his findings in an obscure journal a few years after Darwin’s On the Origin of Species appeared in 1859. It would take nearly a century more before James Watson and Francis Crick deciphered the structure of DNA, the molecule that contains the manual for building an organism. Yet Darwin was still able to describe a mechanism — natural selection — for how evolution shapes life on Earth. That’s like describing how a car works without knowing about the existence of internal combustion engines.



But while Darwin achieved his insights without molecular help, biologists today are intimately familiar with the molecules responsible for the diverse array of plants, animals and other organisms that populate the planet. The study of genes has revealed evolution as essentially a high-stakes poker game in which organisms draw randomly from a deck of genetic choices. At stake is the chance to pass along genes to the next generation. Sometimes the hand is good enough to get ahead in the game, but some hands are losers, perhaps to the extent of extinction. By studying the winners, scientists are learning how the forces of evolution work on DNA, the biochemical repository of an organism’s entire natural history. DNA records the mutations that helped some animals to survive ice ages while others perished, the nips and tucks that make animals more attractive to mates, the big leaps that allowed plants to become domestic crops — they’re all there, written out in a simple alphabet of four letters.
Each organism has its own book of life, but it’s not a just-so story. The genomes of living things are constantly undergoing editing and revision. And each individual has its own edition of its species’s book, shaped by natural selection and the other, perhaps less-appreciated forces of genetic mutation, recombination and drift.
In recent years, the U.S. National Institutes of Health and private companies have sponsored programs to build a library of species’ books, with projects to decode the genomes of humans, chimpanzees, bonobos, dogs, cats, cows, duck-billed platypuses, opossums, orangutans, bacteria, fungi, corn, wheat, bees, fruit flies, worms and a menagerie of other creatures large and small. Comparing the genetic records from these genomes will help researchers piece together a history of how Earth’s current life evolved. But that work has barely begun, and many questions remain to be answered.
Scientists, for example, still don’t know how cells evolved, including how former bacteria came to live inside cells as mitochondria and chloroplasts. (Mitochondria are tiny power plants that supply cells with energy, and chloroplasts are the organelles in plant cells that carry out photosynthesis.) Another mystery is how the complex structure of genes in eukaryotic organisms — in which the genetic material is encased in a nucleus — evolved. Researchers also debate how the shapes and forms of organisms came to look as they do. One of the biggest unanswered questions is whether life on Earth was destined to evolve the way it has.
Limits on evolution
At some time in your life you’ve probably asked yourself if, given a chance to do it all again, you would do it the same way.
Scientists have been asking the same question about evolution, but they’ve been getting different answers.
Play a poker game, rewind it to the beginning, start again and see what happens. Would the game play out the same every time? Stephen Jay Gould, the late evolutionary biologist, didn’t think so. If you replay the game, the shuffled cards will turn up a little different each time, and the order in which the cards are drawn can have profound consequences for the outcome. Replaying the “tape of life” from some point in the past would produce very different life-forms than the ones we have today, Gould thought.
Other scientists disagree. Organisms are dealt a finite number of genes and so must choose from a limited menu of evolutionary options, narrowing the directions the organisms can go in a particular environment. “The evolutionary routes are many, but the destinations are limited,” says Simon Conway Morris, a paleontologist at the University of Cambridge in England. As a result, disparate organisms often end up independently developing the same sorts of structures to solve a particular problem. Take eyes. Although the details of how eyes work vary between species, the basic structures are similar.
But since it’s impossible to turn back time (no matter how easy Superman makes it look) and replay all of evolution again, scientists have devised other ways of investigating the issue.
Richard Lenski, an evolutionary biologist at Michigan State University in East Lansing, is among the scientists hitting the rewind button on evolution. Meter-high letters taped to the windows of his lab spell out the lab’s motto: EVOLVE. In the center of the “O,” the face of Charles Darwin peers out toward the football stadium.
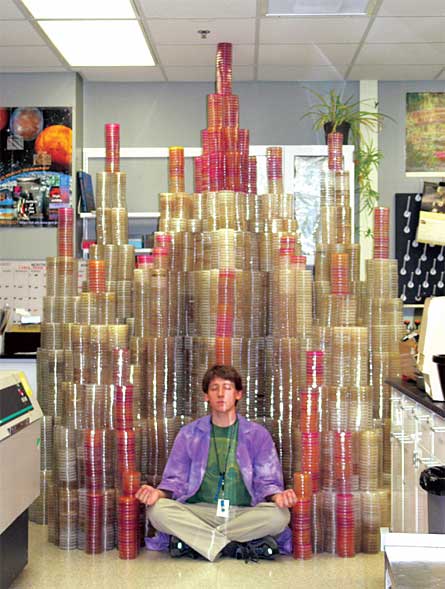
Inside the lab, a dozen glass flasks containing what looks like clear liquid swirl in a temperature-controlled incubator. Although the naked eye can’t see them, millions of E. coli bacteria grow in the flasks, doing what the window exhorts. Lenski started the cultures in 1988, intending to follow the course of natural selection for several hundred generations. Now, 20 years later, the cultures are still growing and have produced more than 45,000 generations of bacteria each.
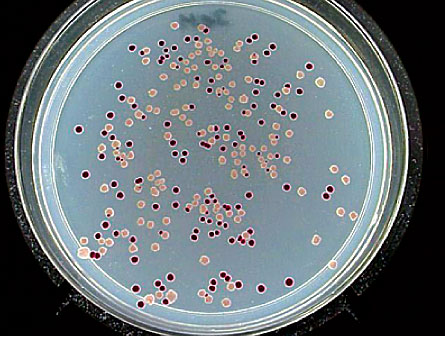
Lenski inoculated each of the 12 flasks with bacteria from the same ancestor, so they all started out with the same genetic deck of cards. Only one gene differed among the bacteria at the beginning — six flasks contain a marker gene that makes the bacteria red when grown on certain media while bacteria in the other six grow white. The marker gene doesn’t affect the strains’ fitness — the term biologists use to describe the capacity of an organism to compete against others to pass on its genes — but it does help researchers distinguish between two lineages of the bacteria during competition experiments.
Each flask contains media with only the minimum requirements for survival — some glucose (a sugar that bacteria use for food) and a few other nutrients. The bacteria replicate, or divide, six or seven times daily, creating a new generation with each round. Each division shuffles the cards and produces genetic changes and mutations, some of which may help or hurt the bacteria’s ability to compete for glucose and win the evolutionary poker game. The next day, a dilution is done, with 10 percent of the culture within a flask transferred to a new flask, and a new hand is played. (Every 500 generations or so, the remaining 90 percent is frozen and stored for later experiments.) The dilution acts as a population bottleneck, randomly selecting a subset of the bacteria (and so a subset of accumulated genetic changes) to continue the experiment.
These 12 flasks “represent the stripped-down bare essentials of evolution,” says Zachary Blount, a graduate student in Lenski’s lab. The environment never changes. No new genes enter the system from migrating microorganisms. And the scientists take no action to affect the course of evolution within the flasks. Only the intrinsic, core processes of evolution influence the outcome, Blount says.
Lenski and his colleagues have watched the game play out, occasionally analyzing DNA to peer over the players’ shoulders and find out what cards they hold. On the surface, the populations in the 12 flasks seem to have traveled similar paths — all have grown larger and become more efficient at using glucose than their ancestors. And many of the strains have accumulated mutations in the same genes. Notably, though, no one strain has developed exactly the same genetic changes as another.
Randomness is an important part of the evolutionary equation, as the experiment illustrates. During the first 2,000 generations, all of the 12 populations rapidly increased in size and fitness. But then cell size changes and reproductive and glucose efficiency gains began to slow down, hitting the evolutionary equivalent of a dieter’s plateau.
After 10,000 generations, it became apparent that not all the flasks were on the same trajectory. Though the cells in all the flasks became larger, each population differed in the maximum size the cells reached. The populations also differed in how much fitter they were than their ancestors, when the researchers grew them in direct competition. The “experiment demonstrates the crucial role of chance events (historical accidents) in adaptive evolution,” Lenski and his colleague Michael Travisano wrote in a 1994 paper.
The experiment has progressed, and several of the flasks now contain mutater strains, bacteria that have defects in their DNA replication system. Such defects make mistakes more likely to happen every time those bacterial strains copy their DNA to divide. Sometimes a mistake can have lethal consequences, damaging a gene crucial for survival. But other times coloring a bit outside the lines creates opportunity for advancement.
Even within a given flask, some bacteria take slightly different paths. One flask now contains two separate strains — one that evolved to make large colonies when grown on petri dishes, and one that makes small colonies. The large- and small-colony strains have coexisted for more than 12,000 generations. The large-colony producers are much better at using glucose so they grow quickly, but they make by-products that the small-colony producers can eat. Each of the populations, both large and small, have improved their ability to use glucose over the generations. By at least one measure, the two populations could constitute separate species, Lenski and his colleagues proposed in 2005 in the Journal of Molecular Evolution.
Still, it seems that Conway Morris was basically right: Though the details were different, replaying evolution in a dozen flasks produced very similar outcomes in each. But then something completely unexpected happened.
After about 31,500 generations, glucose-eating bacteria in one of the flasks suddenly developed the ability to eat a chemical called citrate, something no other E. coli do, the researchers reported last June in the Proceedings of the National Academy of Sciences (SN Online: 6/2/08).
“They’ve been eating the main course for thousands of generations,” Lenski says. “They didn’t realize that there was a dessert tray around the corner.”
The switch was clearly a radical change of destination for the bacteria. The inability to eat citrate is a biochemical hallmark of the E. coli species, so by some definitions, the citrate eaters in that flask are no longer E. coli, but a different species.
But a single change did not a citrate eater make. The researchers found that the bacteria went through a series of steps before evolving the ability to use citrate. One initial mutation happened at least 11,000 generations before the citrate eaters appeared. Lenski and his crew don’t yet know which specific DNA changes led to citrate use, but the researchers have enough evidence to say that the ability to use citrate is dependent, or contingent, upon those earlier changes. And even the bacteria that have undergone those initial changes are still not guaranteed to find the dessert cart. Blount tested 40 trillion bacteria from earlier generations to see if any could evolve the ability to eat citrate. Fewer than one in a trillion could.
The profound difference between the citrate eaters and the other 11 strains, as well as the dependence of the citrate change on earlier mutations, seems to suggest that Gould was also right: Replaying evolution will result in some surprise endings. “The long-term evolution experiment with E. coli provides some of the best evidence for both Conway
Morris and Gould that one could ever hope to see,” Lenski says. “Conway Morris ‘wins’ based on the number of changes that fit his pattern, but Gould might prevail if weighted by the profundity of change. Both perspectives are important contributions, and they are not mutually exclusive.”
Now the researchers are watching to see if citrate-eating bacteria will evolve in other flasks, and if citrate eaters will eventually reject glucose and feast only on citrate. Such a transformation would probably herald the birth of a new species. “It would be amazing,” says Blount. “It would be like teenagers who no longer like to eat pizza — they prefer broccoli.”
When the game changes
While Lenski’s experiment takes place in a constant environment, natural evolution must cope with a messier reality. In Steven Finkel’s lab at the University of Southern California in Los Angeles, a long-term experiment is showing how evolution plays out in a closed and ever-changing environment, more like the real world. This allows Finkel to focus on how evolution and environment are interwoven.
Finkel didn’t start out to test evolution in changing environments. The experiment was prompted by a graduate student’s casual remark that E. coli will live a long time. Finkel asked, “How long?”
“A long time,” the student responded.
“So we set up some experiments to see how long they would live, and they just would never die,” Finkel says. The immortal bacterial cultures are teaching scientists a few things about how organisms change their environments and adapt to changes wrought by outside forces.
Some of the cultures in Finkel’s laboratory have been growing for more than five years. The bacteria started out with the same genetic background, or so the researchers like to tell themselves. In reality, each flask started with a single bacterial colony, containing perhaps 50 billion individuals. Given that DNA replication systems aren’t perfect, one in every 10,000 cells probably carries a typo in at least one letter of its instruction manual. Such DNA typos are known as point mutations.
Finkel and his colleagues placed the bacteria in a rich broth full of sugars and many other nutrients and then just let them grow. After a short initial lag, the bacteria began growing like gangbusters, a phenomenon known to microbiologists as “log phase” because the bacteria increase their numbers logarithmically. Once the nutrients start to run out, the bacteria stop growing so quickly and settle into a senescent state. After a few days, millions of bacteria die, spilling their guts into the surrounding media and providing food for survivors.
It’s the postapocalyptic survivors that interest Finkel. As 99 percent of their comrades die off, the surviving bacteria feed on the carcasses of the dead and on metabolic by-products of other survivors. Thus the bacteria change the environment in which they live. It doesn’t take long for the cultures in each flask to go their own ways. Within a month, the bacteria in the various flasks convert the light honey color of the broth into a spectrum ranging from light amber to dark amber,
Finkel says. And his nose tells him the cultures are different as well. Microscope examinations reveal that the originally rod-shaped bacteria take on a wide variety of shapes; in one flask, some of the cells never cut the apron strings during cell division, forming long strands resembling linked sausages.
Yet as different as the bacterial populations appear, they also have something in common. All of the cells that have gone through the valley of death and come out the other side are tougher than naive bacteria. And the older the cells get, the more competitive they are, so that 20-day–old cells will drive 10-day–old cells to extinction, and 30-day–old cells beat 20-day–olds. Finkel calls that phenomenon “growth advantage in stationary phase,” or GASP.
On the surface it appears that the number of surviving cells stays constant. But underneath, different mutants rise and fall in number, like waves crashing on the beach, Finkel showed in a 2006 review published in Nature Reviews Microbiology.
The ability for older cells to compete better has been traced to mutations in four genes. Three of the genes allow the bacteria to feast more readily on certain amino acids. One of the genes encodes a key protein, RpoS, needed to turn on stress-response genes. The protein gives the green light to turn on genes under certain conditions. When cells are under stress — for bacteria, stress means high salt, low or high temperatures, broth that is too acidic or alkaline, or other environmental extremes — RpoS turns on genes that help the bacteria cope. But the protein is not necessary when cells aren’t under stress. In fact, it takes resources away from the cells’ main “go” signal, RpoD, a protein critical to normal function. Inactivating or handicapping RpoS makes more resources available for other genes.
Many of the GASP cells contain changes in rpoS, the gene for RpoS, but they don’t all have the same change, Finkel and colleagues reported in 2003. Nearly all of the changes reduce activity of RpoS to 1 percent or less of its normal activity but don’t abolish it entirely. Low levels of RpoS are a fixture in bacterial populations that have GASP.
But just because a mutation serves an organism well under some conditions doesn’t mean it’s always beneficial. Thomas Ferenci, a microbiologist at the University of Sydney in Australia, reviewed what happens to rpoS mutants under a variety of environmental conditions in the May 2008 Heredity. Depending on a cell’s genetic background, an rpoS mutation might give the strain a big boost in fitness or make an undetectable difference. And even if the mutations are beneficial under most conditions, the changes hold the bacteria back when the environment changes. If salt concentrations go up, the temperature drops, bacteria lack oxygen or encounter a toxin, then rpoS mutants, less able to cope with certain types of stress, don’t become established members of the community as quickly as they do under other conditions.
Natural selection works for rpoS mutants in some environments and against them in other conditions. “Selection is a deterministic force pushing relentlessly in one direction,” says Michael Lynch, an evolutionary biologist at Indiana University in Bloomington. That direction is toward ever-greater adaptation for the environment in which a population finds itself. But most environments are in a constant state of flux and, as Darwin was careful to point out in his introduction to the Origin of Species, selection isn’t the only evolutionary force at work.
Sex, chance and genes
Random genetic drift is an evolutionary force to be reckoned with too. And, as with selection, molecular biologists are helping to reveal its workings.
Drift by any other name would be known as chance. The number of individuals that carry a specific genetic variation within a population — what scientists call the frequency of a gene variant — can change at random, bobbing along like driftwood on the ocean. The indiscriminate nature of drift doesn’t always work to organisms’ betterment.
“Drift doesn’t care about fitness,” Lynch says.
Drift can haphazardly make a detrimental gene prominent in a population, or accidentally eliminate beneficial mutations — especially in small populations. Imagine two versions, or alleles, of a gene as the head and tail of a coin. Every time an organism reproduces, the coin is tossed to see which allele will be passed to the offspring. In a large population, coins will flip many times and the number of heads and tails will be roughly equal. But in a small population, runs of heads or tails can skew the outcome in favor of one or the other allele, maybe even eliminating one version altogether.
That’s a simple example of what drift can do, but Lynch thinks it also accounts for some complex traits, such as the complicated structure of genes in eukaryotic organisms — including multicellular beings like people and plants and unicellular life such as yeasts.
Drift causes noise in the evolutionary process, says Lynch. But there is yet another force that mixes things up — genetic recombination. Recombination is an essential element of sexual reproduction. In general, each parent contributes a single copy of each chromosome to its offspring. Before mom and dad divvy up the genetic goodies to hand down to the children, the two copies of each chromosome are lined up and matched like pairs of socks. When the two chromosomes are zipped together, they swap chunks of DNA, giving each egg or sperm a different combination of the parents’ genes.
Genes follow each other along a chromosome like freight cars on a train follow the lead engine, unless recombination happens. So if one gene develops a lethal mutation it may doom the other genes on the chromosome, like a train car that gets unbalanced and derails the train. Similarly, a beneficial mutation might get trapped on a slow train to nowhere if not for recombination shuffling the mutated gene’s position on the chromosome. Or a particularly good mutation may create such a powerful engine that natural selection can’t resist taking along whatever’s attached — like an engine dragging a decrepit train.
Low rates of recombination enhance the effect of drift because “beneficial alleles could get trapped in bad backgrounds,” Lynch says. Natural selection would derail some trains, taking “good” genes along with the bad. “That’s simply because you are a victim of the surrounding genetic material,” he says.
Recombination allows the cars to uncouple and switch around, creating faster, more efficient trains. Once removed from a bad neighborhood and pasted in a beneficial or neutral stretch of chromosome, an allele’s attributes can shine, and natural selection can act on the allele without any worries about the company it keeps. In this way, the process helps to increase the efficiency of natural selection, Lynch says.
Researchers are still debating the details of how selection works together with mutation, recombination and drift to shape genes and help organisms adapt to their environments, producing the abundance of species around today.
“We’re peeling back the onion of the evolutionary process,” says Sean Carroll, a developmental and evolutionary biologist at the University of Wisconsin–Madison. “The question is no longer ‘does adaptation happen?’ but ‘how does it happen?’ ”
Evolutionary tweaks
In the past few years, scientists have learned that genes work together in vast networks to regulate every process in the body. Of special interest to many evolutionary biologists are transcription factors, proteins that are important for controlling the timing and placement of gene activity during development (and later). Each transcription factor may govern multiple genes, even hundreds of genes.
“We’re into scores of direct targets,” Carroll says. “More than we imagined. More than we even have an explanation for.”
Implications of such vast gene regulatory networks are clear for Carroll. Altering the structure of a transcription factor to better regulate one gene could have effects on hundreds of other genes. Tinkering with a transcription factor doesn’t just alter the shape of a fin, add a horn or move a spine. No. These molecules are so important and work in so many different parts of an organism that changing the transcription factor itself is likely to affect nearly everything about the living thing. Most of the time, such far-reaching changes are bad, even lethal. Carroll and others have been working to understand how body shapes and coloring morph in animals. If changing transcription factors could be catastrophic, organisms must make molecular tweaks elsewhere to create a new look.
Carroll is a leading proponent of an idea called the cis-regulatory theory. (Cis refers to a region adjacent to the gene or on the same chromosome.) The theory holds that altering the control region of a gene to change some feature of an animal or plant would produce fewer side effects than tinkering with the proteins that direct construction of the features. So an organism can change one part of its body without affecting the rest simply by adding a few more switches and buttons to its control panel (or taking some away), or by rewiring a switch to work at a different time or govern development in a new location.
Fish called three-spined sticklebacks have provided some of the most direct evidence that the cis-regulatory theory could be correct. The fish live in saltwater but swim into rivers to spawn. That habit led to isolation of many of the fish in inland freshwater lakes at the end of the last ice age. Over the past 10,000 years the fish have adapted to their new homes, says David Kingsley, an evolutionary biologist at Stanford University.
In the ocean, sticklebacks wear armor and sport pelvic spines that protect them from sliding down the throats of predators. Fish stranded in freshwater lakes found themselves without the fishy predators they knew in the ocean, but some encountered deadly insects, such as dragonflies.
Dragonflies grab the sticklebacks by their pelvic spines and eat the fish sideways, so the feature that once offered protection became a liability. Over time, some populations of fish have shed their armor and their pelvic spines.
Kingsley and his colleagues discovered that a protein called PITX1 is responsible for building the pelvic spines. The protein is made in the hind limbs of many different animals, including humans. A group of researchers from Washington University in St. Louis showed that a mutation in PITX1 in humans caused clubfoot in members of a family. The team published the research in the Nov. 7 American Journal of Human Genetics. The protein also controls development of the pituitary gland and facial development. Defects in PITX1 can lead to cleft palate.
But when Kingsley and his colleagues examined the gene encoding PITX1 in stickleback fish with and without spines, the researchers found no differences. That led them to believe that the defect must lie in the control panel for PITX1 and not in the gene itself. But the scientists had no direct evidence that changes in the control panel were responsible for the missing spines.
Recently, Kingsley’s group did discover that some of the stickleback species that have lost spines also lost a portion of the control panel that turns PITX1 on in the pelvis. Restoring the lost switch also restored spines, Kingsley told science journalists gathered in Palo Alto, Calif., in October at a conference sponsored by the Council for the Advancement of Science Writing.
“That’s an ‘i’ that has needed to be dotted,” says John “Trey” Fondon III, an evolutionary biologist at the University of Texas at Arlington. “We’ve had some really great circumstantial evidence for cis-regulatory evolution, but the data have been lacking. It’s been a little, what we call, ‘empirically challenged.’”
Fish aren’t the only animals providing evidence for how evolution works in genes. Fondon and others have turned to man’s best friend to figure out how genes influence body shape and size, behavior and other traits. Dogs come in an astounding number of variations, with the smallest dogs, Chihuahuas, weighing under six pounds and the largest breeds weighing more than 100 pounds.
A group of scientists from the National Institutes of Health and collaborators traced body size in dogs to a variation of the insulin-like growth factor gene (IGF1). Within the gene itself, researchers found no differences between large dogs and small dogs. But dog breeds that weigh less than 20 pounds had a common change in the IGF1 control panel, altering how much of the protein is produced, the researchers reported in 2007.
Kingsley cites the IGF1 finding as further evidence that changes in control regions can account for surprisingly large differences in body shapes and sizes. But Fondon says he isn’t ready to swallow the “cis-regulatory Kool-Aid” just yet.
In contrast to the cis-regulatory theory, Fondon and his colleagues have found evidence that tampering with transcription factors can change specific features without having disastrous consequences for the whole organism. The team focused on repeats of amino acids within proteins. Proteins work a bit like Swiss Army knives with various tools tied together in a single package. The repeated amino acids are often found between the stretches of amino acids that form each of the tools. Fondon reasoned that slightly altering the number of repeated amino acids, each of which are encoded by repeated three-letter DNA sequences, might subtly change the function of the protein, creating a variant that could be put to an evolutionary test.
But Fondon realized that hypothesis also had its weak point: Repeats in DNA tend to mutate at high rates. The machinery that replicates DNA loses its place when reading the same letters over and over and over again. Sometimes it slips up and skips a repeat or adds an extra. High mutation rates can be dangerous because of a higher chance of catastrophic error. In some families, extra DNA triplet repeats in the gene for the huntingtin protein can lead to Huntington’s disease. Often the number of repeats grows with each generation, causing people to develop the disease at younger and younger ages.
“I thought selection wouldn’t tolerate this kind of crap in our genes,” Fondon says.
But when he created a computer program to find genes in dogs that contain such repeats, he found a surprising number. “The top half of the list was a who’s who of development,” he says. This list includes genes that control bone development and the homeobox genes, which encode transcription factors that direct construction of an animal’s body, distinguishing head from tail and back from front, and guiding the positions of limbs and appendages. These genes are found in almost every type of animal on Earth, from sponges to people. Even fungi and plants have some forms of homeobox genes.
Expanding and contracting the number of amino acid repeats in certain homeobox genes seems to give dog breeds some of their distinguishing characteristics.
“If what the protein does is a verb, a repeat is an adverb,” says Fondon. The repeats don’t change what the protein does; they just make it happen more quickly, slowly or frequently.
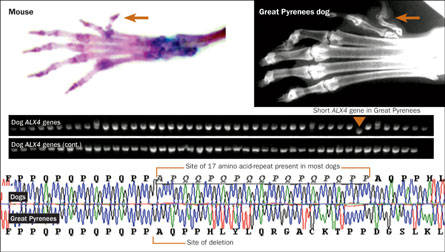
For instance, in Great Pyrenees, deletion of a repeat in the ALX4 gene leads to the formation of an extra toe on the back feet, a hallmark characteristic of the breed. Mice with defects in ALX4 also grew extra digits on their back feet.
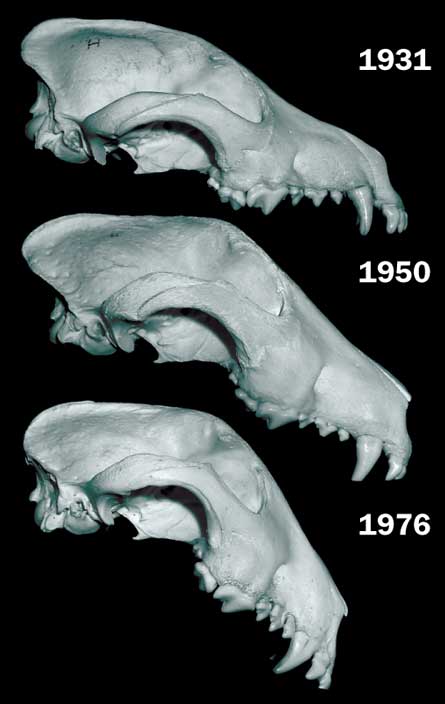
A protein called RUNX2 governs genes that help control facial development in dogs. Fondon and his colleagues found that changing numbers of repeats within RUNX2 are associated with ongoing exaggeration of certain face traits. Scientists have documented such changes in the bull terrier’s RUNX2 protein between 1931 and today. Modern bull terriers have fewer repeats of a certain amino acid sequence than members of the breed did in 1931. That doesn’t sound like a big change, but could be one factor contributing to the flatter faces seen in today’s bull terriers.
Carroll doesn’t deny that mutations happen in transcription factors too, but he thinks such mutations probably affect genes that play a more limited role in development. Regulatory changes are more likely in genes that govern development of many different parts of the body, he says.
But others don’t think it has to be all regulatory mutations or all protein changes that create novel traits in animals. Vincent Lynch, an evolutionary biologist at Yale University, and his colleagues discovered that both types of changes were needed for pregnancy to evolve in mammals.
Control issues
When most female mammals get pregnant, the embryo attaches to the wall of the uterus. Normally something burrowing into the body causes the immune system to take up arms and oust the intruder, but that would make pregnancy impossible. So placental mammals turn up production of prolactin, a protein that calms the immune system and does other jobs that allow an embryo to develop safely.
Yale’s Lynch and his colleagues discovered that the evolution of pregnancy probably happened in several steps. The first step was that a jumping gene, called a transposon, hopped into the control panel of the prolactin gene. The transposon brought with it a switch operated by the homeobox protein HOXA11. Over time, HOXA11 developed changes that allowed it to work with other proteins to more precisely control prolactin production, Lynch and his colleagues reported in the Sept. 30 Proceedings of the National Academy of Sciences. Only HOXA11 from mammals turns on prolactin, the researchers showed. HOXA11 from chickens, platypuses and opossums (all animals without a placenta) failed to turn on production of the pregnancy-associated protein.
Such complementary changes to proteins and their genetic control panels help evolve a toolkit that organisms can use for every occasion, Lynch says.
“A hammer in a toolbox can be a ball-peen hammer. It can be a hammer that pulls nails. It can be a mallet, but it’s still a hammer. It evolves to its own context,” he says.
Animals, plants, bacteria, archaea, fungi and all organisms on Earth evolve to their own contexts as well. Scientists are now beginning to learn how tweaks and major changes on the molecular level enable adaptation to environments. The picture is painted in DNA, but it’s far from a completed masterpiece. Changing environments coupled with the forces of natural selection, mutation, recombination and drift are continually reworking the painting. Only time will tell how the landscape will morph — and its inhabitants with it.