Probing Ocean Depths: Photosynthetic bacteria bare their DNA
Scientists call it the invisible forest, the immense mass of ocean-dwelling micro-organisms–algae, bacteria, and plants known as phytoplankton–that perform photosynthesis as trees do. Converting light, water, and carbon dioxide into energy, these microbes produce nearly the same amount of oxygen worldwide as land plants do and influence the climate by sequestering carbon inside the oceans.
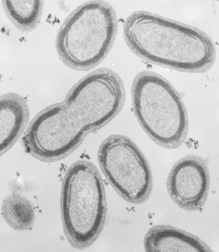

Biologists have now deciphered the full DNA sequences, or genomes, of two kinds of cyanobacteria at the heart of this ocean forest. “They are the most abundant photosynthetic cells on the planet,” says Sallie Chisholm of the Massachusetts Institute of Technology.
“About half the oxygen you breathe is produced by the oceans, and a very significant portion of this oceanic production comes down to these cyanobacteria,” adds Nicholas H. Mann of the University of Warwick in Coventry, England.
The photosynthetic bacteria targeted by the new research are known as Prochlorococcus and Synechococcus. The former includes strains that thrive in the top 100 meters of the ocean and other strains, adapted to less light, that primarily populate depths of 100 to 200 m. Synechococcus tends to inhabit the top 20 m of ocean water.
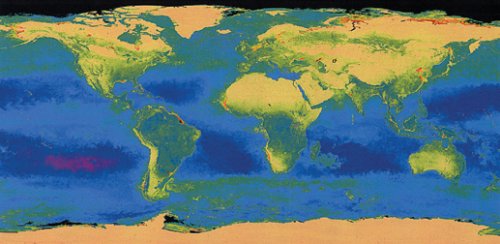
GREEN WATERS. Photosynthesis depends on chlorophyll, and this map shows the amount of chlorophyll in the oceans. Midocean purple and blue represent low ocean-chlorophyll concentrations, while green, yellow, and red in coastal waters indicate progressively higher amounts. |
The analyses of three Prochlorococcus strains, to be reported in the Aug. 19 Proceedings of the National Academy of Sciences and an upcoming Nature, were spearheaded by Gabrielle Rocap of the University of Washington in Seattle and Alexis Dufresne of the University of Paris. Rocap’s group, for example, found that a low-light Prochlorococcus had nearly 2,300 genes, while a shallower-dwelling form had about 1,700 genes.
Gene differences between the strains could explain why they thrive at diverse ocean depths, but each one’s gene number itself may not be telling. Chisholm, who worked with Rocap, notes that the third Prochlorococcus strain, studied by Dufresne’s team, is adapted to low light but has fewer than 1,900 genes.
In part to resolve a mystery, Brian Palenik of the Scripps Institution of Oceanography in La Jolla, Calif., and his colleagues studied the genes of a strain of Synechococcus. “It has a very unique form of motility. It’s able to swim, and no one knows how,” he says.
His group’s genome analysis, also in an upcoming Nature, confirms that the bacterium lacks most genes crucial to known modes of swimming. The researchers did identify a novel gene that might contribute to the strain’s motility. The cyanobacterium can’t swim if this gene, encoding one of the largest bacterial proteins ever documented, is mutated.
Unexpectedly, the microbe also contains genes for proteins that pump toxic substances out of a cell. That suggests the cyanobacterium, even in the open expanse of an ocean, regularly encounters other microbes that seek to kill it with chemical weapons, says Palenik.
Furthermore, he and his colleagues found evidence in the cyanobacterium’s DNA of past infections by viruses known as bacteriophages or phages. Scientists have recently realized that such viruses have infected many ocean bacteria, providing one way that the bacteria acquired new genes (SN: 7/12/03, p. 26: All the World’s a Phage).
Many phages, however, don’t make a permanent home inside their host. In another Nature report, for example, Mann and his colleagues reveal the DNA sequence of a virus that infects and replicates inside Synechococcus strains, eventually killing them. Mann’s team discovered that this particular phage contains two genes for proteins required by the Synechococcus‘ photosynthetic process.
Normally, sunlight damages these two proteins, so bacteria must keep making replacement copies. But Mann suspects that when the virus infects a Synechococcus, the phage turns off most of the bacterium’s genes. So, to keep energy flowing for its own reproduction, the phage supplies copies of the photosynthetic genes. “It’s not an act of altruism on the part of the phage. It’s a cynical takeover of the cell,” concludes Mann.
One theme emerging from these studies of cyanobacteria, notes Donald A. Bryant of Pennsylvania State University in State College, is that the organisms don’t have as many genes devoted to sensing and reacting to the environment as many other bacteria do. “The ocean is a big place and relatively constant as an environment. The concentration of many things in the ocean is relatively fixed, so there’s no reason to sense change,” he says.
Iron concentrations, however, do change in the ocean, and there’s often a scarcity of that metal. Perhaps as a result, the cyanobacteria possess extra genes for enzymes that require nickel and copper, rather than iron, for their function. “That’s a particularly interesting adaptation, something rather novel,” says Bryant
****************
If you have a comment on this article that you would like considered for publication in Science News, send it to editors@sciencenews.org. Please include your name and location.
To subscribe to Science News (print), go to https://www.kable.com/pub/scnw/
subServices.asp.
To sign up for the free weekly e-LETTER from Science News, go to http://www.sciencenews.org/subscribe_form.asp.