DOCTORING DNA’S DIRECTIONS The longfin inshore squid edits more than half of the RNA that codes for proteins in its nervous system.
Roger Hanlon, Marine Biological Laboratory, Woods Hole, MA
A species of squid can alter vast amounts of its genetic instructions after they are copied from DNA into RNA, a study reported January 8 in eLife finds. The squid edit their RNA more than any other species yet studied, allowing them to create proteins that are not coded for in their DNA.
“If you edit RNA at multiple sites, you really start to create diversity,” says study coauthor Joshua Rosenthal, a neurobiologist at the University of Puerto Rico. “From a single gene you can make a lot more proteins.” The researchers have not determined what role the edited proteins play, but speculate that the changes may help the animals adapt to their surroundings.
The researchers compared squid DNA to a transcript of the animals’ protein-coding RNA. So much of the RNA had been altered that, the authors argue, RNA editing may be a common process in the animal kingdom to modify biology’s central dogma, which describes a more direct transition from DNA genetic information to RNA to proteins.
“We have this central dogma where information flows from the genome to the RNA to the protein,” says physicist Eli Eisenberg of Tel Aviv University, another coauthor. “RNA editing is already an exception to this paradigm.” But, he says, “it was considered to be a rare exception.”
When a cell builds proteins, it copies the instructions encoded in DNA into RNA. The RNA ferries the information to parts of the cell where it can be translated into proteins. Editing occurs when the genetic instructions are changed after they have already been baked into RNA.
The most frequent target of editing is adenosine, one of the four major building blocks of RNA. An enzyme can transform adenosine into a compound called inosine. The cell reads inosine as if it were another RNA building block, guanosine.
Most RNA editing in humans occurs in parts of the RNA that get spliced out or are left untranslated when proteins are made; it does not change the proteins that are coded.
RNA editing has only been investigated in a few species whose genetic instruction manuals, or genomes, have already been compiled. “Everyone up to this point used a genome as a point of reference,” says Rosenthal. “It’s like an anchor for their analysis.”
However, researchers have not yet analyzed the entire genome of the longfin inshore squid (Doryteuthis pealeii). Eisenberg’s team got around this obstacle by creating a transcript of all the squid RNA that codes for proteins. The team matched DNA from the squid to its corresponding areas in the RNA, then looked for discrepancies between the DNA and RNA.
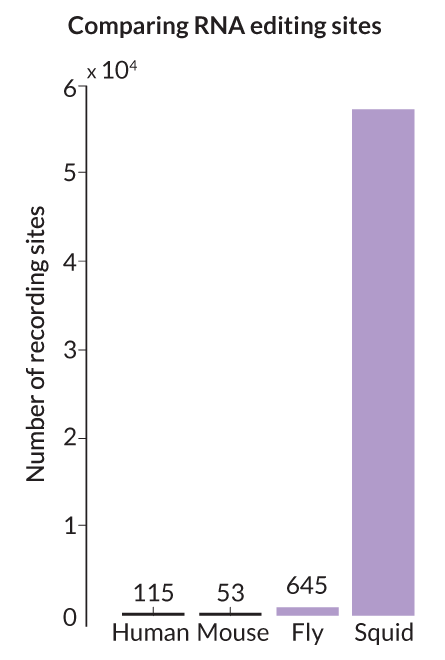
Most of the altered RNA appeared in nervous system tissue. More than 57,000 sites — nearly 60 percent of the RNA in these areas — were edited, orders of magnitude more than in any other species studied so far.
The team compared the number of incongruities with those in human and rhesus macaque transcripts. “The squid gave us this huge spike,” says Rosenthal. “The humans and monkeys don’t show that.”
Jin Billy Li, a geneticist at Stanford University, looks forward to future research from the team to find out why the squid have so many edited spots in their RNA. “This group has done a wonderful job to computationally identify … editing sites without a reference genome,” he says. “It’s really surprising to see so many events.”
In the future, the researchers want to find out why the squid have so much edited RNA and whether this phenomenon shows up in other members of the animal kingdom. They also plan to investigate whether the RNA editing always results in proteins that are beneficial for the squid, and if the modified proteins allow the animals to adapt to changes in their environments.