Alphabet of Life
Searching for clues to the genetic code's origin
“Omg. u no how 2 do the bio hw?”

Texting uses a peculiar alphabet. It keeps messages brief but still encodes enough meaning for students to communicate about homework, coffee dates and crushes — all while accommodating the occasional typo.
The genetic alphabet, the letters used as the blueprint for all life, balances brevity and clarity in a similar way. Just four letters combine to spell out the more than five dozen three-letter words that encrypt the information needed to make all the cells in the human body, and any other body as well.
Figuring out how life’s code came to be is nature’s original homework problem, and it isn’t easy: It’s like studying how people in Paris talk today to determine what the first Latin alphabet must have been. Attempts at deciphering the code’s origins are also complicated by the fact that, unless another iteration of life turns up, say on a distant planet, scientists have only one version to study.
“We only have one experiment, and it’s extremely hard to repeat this experiment,” says physicist Tsvi Tlusty of the Weizmann Institute of Science in Rehovot, Israel.
Without a way to replicate life’s earliest days, coming up with a theory to explain the code is like interpreting a Rorschach test. But that doesn’t stop scientists from trying. Some are now finding that outside pressures, such as a need to minimize error, may have driven the code to evolve the same way as texting — through en masse trial and error. Chemical attractions between molecules, others report, could have set the code’s destiny.
“These discoveries make the whole field legitimate rather than a matter of pure speculation,” says molecular biologist Eugene Koonin of the National Center for Biotechnology Information in Bethesda, Md.
A shared code
All life on Earth — people, pandas, tulips and slime molds — uses the same genetic letters in its DNA. These letters — A, C, T and G — stand for DNA’s four bases (adenine, cytosine, thymine and guanine). With sugar and a phosphate, these molecules form the basic units of DNA, called nucleotides. And the letters provide instructions for making proteins, essential cellular players that kick-start chemical reactions, serve as scaffolding and act as messengers.
To make a protein, the two strands of the double helix–shaped DNA unwind, and a complementary strand called messenger RNA forms alongside one of the exposed arms of the helix. Messenger RNA is a copy of DNA with chemically similar letters (though messenger RNA has a U for uracil instead of thymine). This RNA is moved to a cellular factory called a ribosome where, through a process called translation, the RNA gets decoded and proteins are constructed.
Every three nucleotides in the messenger RNA spell a genetic word, called a codon, that codes for a specific amino acid. In the ribosome, amino acids are linked to form protein molecules, the way words come together into sentences. The codon for the amino acid methionine often serves as a start signal, while three other codons serve as punctuation marks, telling protein construction to “stop.”
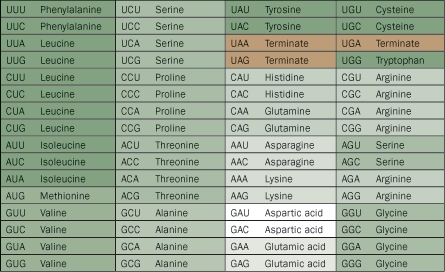
With an alphabet of four letters, 64 three-letter codons are possible. Yet cells make their proteins from only 20 amino acids.
Francis Crick, codiscoverer of the DNA double helix, proposed in 1968 that the code for these 20 amino acids was a “frozen accident,” working well enough to get passed down through generations like an old tradition. If so, on a second go-round life could look completely different, and any life found elsewhere in the universe would probably be unfamiliar as well. But some of the alphabet’s features seem too good to be an accident, so scientists have tried to find logic beyond pure luck.
Some of the words, for example, act like synonyms. Three or four codons, usually identical except for one letter, can stand for the same amino acid, just like hi and hey both mean “hello.” This feature can protect cells from errors: If messenger RNA’s CGA mistakenly becomes CGU, for example, the cell still selects the same amino acid (in this case, arginine).
But even if a mistranslation leads to the wrong amino acid, the code is arranged so that the product will be chemically similar to the intended one. This logic isn’t generally a rule in English: You’d have trouble doing your homework with a “hen” if you really needed a “pen.”
At the same time, the 20 words are different enough that combining them can make bats, bees, birds and bacteria. If the amino acids are LEGOs and the task is to build things for 4 billion years, you had better have a diverse set, with rectangular blocks, joints and wheels, says astrobiologist Stephen Freeland of the University of Hawaii at Manoa.
There are plenty of other ways to arrange the code. By the calculations of a team including Peter Clote, now of Boston College but formerly of Ludwig-Maximilians-Universität München in Germany, there are 1084 alternate codes that assign at least one codon to each of the 20 amino acids (and “stop”). So it would be weird if, by complete chance, the code had such clever traits.
Dodging errors
Tlusty thinks that if the genetic code is so good at what it does, it’s because it adapted under evolutionary pressure, the way the beaks of some of Darwin’s finches developed to be hefty seed crackers or long flower probes depending on available food.
Three pressures, Tlusty argues, could have shaped the current genetic code. First, typos can’t be disastrous — if a random mutation changes one of the letters, the cell should still infer which spelling was intended. Second, the language must be able to spell words with diverse meanings. Third, the language shouldn’t take a lot of resources to write, forcing the cell to make tons of extra molecules.
Assuming an evolving code, and assuming four letters in the alphabet and three-letter words, Tlusty tried to figure out an ideal number of amino acids. He imagined the code as mapping onto a many-dimensioned doughnut of sorts, with all 64 possible codons spaced out so that those that could easily be confused with each other are neighbors. Then he tried to find how many colors, or amino acids, are needed to make a map that obeys his three demands. In this way, Tlusty rephrased a biological question (How many amino acids would a changeable code settle on?) as a classic mathematical one: What is the fewest number of colors necessary to color geographical divisions on a map without any colors touching themselves?
On a two-dimensional map, like a map of the United States, the answer is four colors. In the higher dimensions necessary to map the code, the range of colors is between 20 and 25, Tlusty reported in Physics of Life Reviews in September. That’s spot on for how many amino acids are in today’s code.
Tlusty’s findings support his idea that a changing code could have settled on an optimal number of amino acids. Another team suggests that an earlier code, one that preceded today’s, could have been a superstar at one of Tlusty’s three demands.
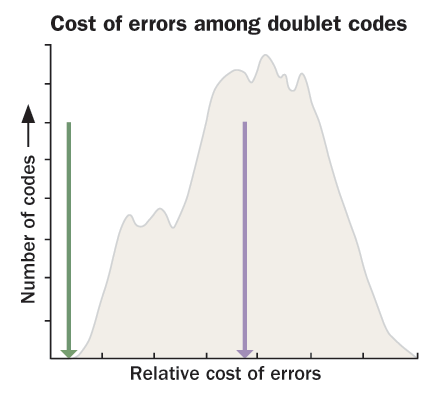
If randomly generated codes are placed on a landscape, with higher elevations designated for codes that are better at preventing errors in protein manufacture, life’s code would be found on the side of an unassuming hill. But a previous code could have been atop one of the highest peaks, Koonin argues.
Because changing the third letter of a codon doesn’t drastically change the amino acid, Koonin and others think that an early genetic code relied on only the first two of its three letters. Such a code could have expressed at most 16 amino acids. Koonin’s team backtracked the code to its most plausible two-letter origins, playing with different codon assignments. In Biology Direct in 2009, the researchers reported that several such codes were exceptionally robust against translation errors, suggesting that minimizing error drove the code’s development early on. When having more than 16 amino acids was advantageous (allowing for more types of proteins), the code started to use the third letter — getting a little worse at avoiding errors.
Chemistry behind the code
While agreeing that there is nothing accidental about the code, other scientists suspect chemistry was a more important driver. They are turning to experiments in modern labs to try to determine which amino acids came first.
In a legendary spark-tube experiment in 1953, Stanley Miller created a handful of life’s amino acids by electrically zapping a chamber filled with hydrogen, water, methane and ammonia gases. And similar follow-up experiments have yielded even more amino acids. A meteorite that crashed in Australia in 1969 contained some of the same amino acids, suggesting that they were forged somewhere in the solar system.
Five amino acids made in spark-tube experiments and found within meteorites — glycine, alanine, aspartic acid, glutamic acid and valine — appear to be related. Each of their codons begins with a G, suggesting that whatever word coded for the first amino acid, G may have grabbed the first position.
“It’s sort of a hand-waving argument,” says Paul Higgs, a bioinformaticist at McMaster University in Hamilton, Canada. “I don’t know if you can really prove that.” But in the absence of a detailed chronology, deep hunches have led Higgs and others to create models that begin with these five amino acids. In 2009 in Biology Direct, Higgs set out a plan for how the remaining amino acids would be added after the first handful were fixed.
Others argue, though, that the first amino acids weren’t the most abundant, but instead were the ones with a natural chemical attraction to RNA, which some scientists think got life off the ground (SN: 7/3/10, p. 22).
Today, every protein found in the cells of every organism has to originate in a DNA or RNA blueprint, as far as biologists can tell. But making DNA and RNA — and proteins — requires proteins. This chicken-and-egg conundrum has flummoxed scientists for decades. In the 1980s, researchers discovered a particular type of RNA called a ribozyme that may be capable of making itself by catalyzing its own synthesis. Many believe the ribozyme could be the chicken and the egg — boosting the popularity of what has been known as the “RNA world hypothesis.”
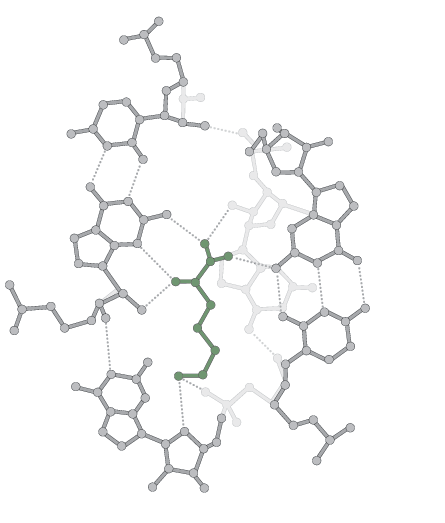
Working in this context, biochemist Michael Yarus of the University of Colorado at Boulder imagines different amino acids being chemically attracted to different strands of RNA.
To test the idea, Yarus and colleagues mounted eight amino acids into test tubes and washed the molecules with a solution containing many RNA snippets. Though the interactions in general are weak, Yarus found that many amino acids have natural docking bays for the sequences of three nucleotides that encode them today. He and colleagues estimated in 2009 in the Journal of Molecular Evolution that there is a one in 1044 chance that the triplets occur at binding sites by pure chance. He suspects that about three-fourths of amino acids currently in the code entered via chemical attraction.
“If you think the triplets were not involved in binding amino acids, then you have to argue that this is all some mistake, some joke nature is playing,” Yarus says.
In his chemical theory, it would be inevitable that certain letters end up coding certain amino acids, like how the word for a cow’s vocalization is moo because that is what the cow sounds like. Any life evolving under similar conditions — say on an extrasolar planet — would thus have a similar code.
“My prediction is if life’s out there, it has a similar core,” Yarus says. “That will be a great day when we find out.”
In November, Wentao Ma of Wuhan University in China proposed a model in Biology Direct for how to get from Yarus’ simple bonding scheme to today’s sophisticated reality. But the proposal now needs evidence to back it up, and some researchers point out that no one has been able to make the ribozyme, a key player in the RNA world, grab the specific amino acids that match up with its nucleotides. Instead, the ribozyme makes up random definitions for genetic words, like in the game Balderdash.
Freeland thinks, because of these major roadblocks, the RNA world may have already passed its prime. “The concept is simply we don’t know anything that can make RNA and use it as a living system,” Freeland says.
He says it’s just as believable that some earlier organism, not directly related to current life on Earth, invented RNA. In that case, scientists would have to go back even farther in time to make sense of the code.
Paying attention to proteins
Going back in time is exactly what researchers at the Georgia Institute of Technology in Atlanta are trying to do in two new approaches.
Biochemist Loren Williams has turned to the ribosome, that cellular factory where proteins are made. Scientists know that some RNA in the ribosome is very densely connected to other RNA through hydrogen bonds, suggesting those RNAs are among the oldest — the way people who have been on Facebook awhile have more friends than those who just joined. By looking in regions with dense connections, Williams has found what he argues is the oldest protein inside the ribosome. The protein’s tail is made of glycine and alanine, Williams and his colleagues recently found, leading them to think that these amino acids may have been the first to join the code.
A second approach, which involves resurrecting ancient proteins, may offer a good test for theories about the code’s origins. Astrobiologist Eric Gaucher, also of Georgia Tech, and his colleagues are comparing individual proteins in different living organisms to try to estimate a most likely protein ancestor, going back billions of years. Such ancient molecules can provide clues about which amino acids early life used, and the team’s findings agree with those from lab experiments and meteorite evidence.
Such ancestor proteins can also provide a valuable check on theories for the early genetic code, says Freeland. If the earliest proteins are made of five amino acids with codons that start with the letter G, for example, then any proposed earliest code had better be able to make those proteins. If it can’t, then that code should be scrapped.
Though not the inkblot it was four decades ago, the code still holds many mysteries. Until they are resolved, some scientists believe, the frozen accident is still a plausible possibility and life on other planets may turn out to be completely different from life on Earth. If texting teenagers could offer any consolation to those in search of answers, it would be “gud luk.”