Blood Work
Scientists seek to identify all the proteins in plasma
In his 1998 book Blood: An Epic History of Medicine and Commerce, author Douglas Starr traced the rise of blood as a commercially exploited tissue. In the preface, he compared blood to oil and suggested that the former is more valuable. At the time of the book, a barrel of crude oil sold for about $13, whereas Starr estimated that the same quantity of whole blood would fetch $20,000–and more than $67,000 if it were processed. “Just like the oil industry, the blood trade involves collecting a liquid resource, breaking it into components and selling the product globally,” he wrote.

Consider the intense demand for plasma, the clear liquid portion of blood that contains therapeutic molecules such as clotting factors and albumin, a protein regularly used to treat shock and other conditions. The companies selling these natural compounds process more than 22 million liters of plasma each year, according to the Plasma Protein Therapeutics Association in Annapolis, Md.
The annual worldwide revenue from plasma-derived therapeutic products is around $5 billion, according to one estimate cited in Starr’s book.
Plasma proteins also provide a wealth of diagnostic tools. For instance, physicians look at high blood concentrations of a protein called prostate-specific antigen as an indication of prostate cancer. And people view their blood concentrations of HDL and LDL, two cholesterol-carrying proteins, as signs of the health of their heart.
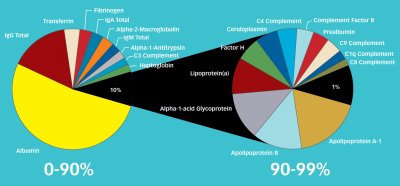
MOLECULAR MIX. Relative abundance of the proteins that make up 99 percent of the protein mass in blood plasma. The final 1 percent represents thousands of proteins. |
Still, there may be a lot more value hidden within plasma. To date, scientists have identified only several hundred of the estimated thousands of proteins typically coursing through a person’s bloodstream.
Between 1977 and 2001, scientists screening human plasma for new proteins added only about 20 proteins to the tally, says N. Leigh Anderson, who founded the Plasma Proteome Institute, a nonprofit research organization in Washington, D.C. “That’s just astoundingly, amazingly bad,” he says.
Indeed, Anderson and other scientists suggest that human plasma contains some of every protein produced by the human body, perhaps hundreds of thousands of different molecules. Identifying this repertoire of plasma proteins could offer new ways to detect–or even treat and prevent–many diseases.
“It’s a very broad and bold opportunity,” says Gilbert Omenn of the University of Michigan in Ann Arbor, who leads an international consortium of scientists seeking to characterize plasma proteins.
The search for plasma proteins is speeding up. Last December, a team at Pacific Northwest National Laboratory in Richland, Wash., reported that it had identified almost 500 proteins in a sample of human plasma. That nearly doubles the number previously known, says team leader Joel G. Pounds.
“We’re right at the cusp of a huge revolution in the field,” says Anderson.
Too much of a good thing
Spin a vial of blood in a centrifuge, and the red and white blood cells, as well as the disklike platelets responsible for normal clotting, sink to the bottom. A clear liquid remains above them. That fluid, the plasma, consists of water with salts, hormones, enzymes, antibodies, and other proteins dissolved in it.
Over the past few decades, Anderson and other scientists have largely used a process called gel electrophoresis to identify plasma proteins. In this approach, investigators put plasma at one end of a slab of gel and run an electric current through it. Migrating with the current through the gel at speeds based on their molecular weight and electric charge, plasma proteins separate from each other.
The researchers then extract purified proteins from the gel.
In a review of plasma-protein research published in the November 2002 Molecular and Cellular Proteomics, Anderson and his father, Norman G. Anderson, estimate that about 290 plasma proteins have been identified in this way. They argue that gel electrophoresis used by itself will find few additional blood proteins.
There’s no doubt that many more proteins lurk in serum, say the researchers. Blood comes into contact with almost all tissues in the body, and damaged or dying cells regularly dump their contents into the bloodstream. An important example is the protein creatine-kinase MB, which some dying heart cells release. Physicians use the protein’s presence in plasma to diagnose heart attacks, and its concentration reflects the amount of heart tissue that has been damaged.
Plasma “is a great place to look for proteins that leak out of tissues,” says N. Leigh Anderson. “The circulatory system is like the canals of Venice. It transports all the good things, and it also transports a lot of junk.”
But it’s not easy to see many of the good things or the junk in plasma because they’re often obscured by a few overwhelmingly abundant proteins. The most plentiful is albumin, which transports several other proteins and maintains blood pressure.
By itself, albumin makes up about 60 percent of the serum-protein content, says Pounds. The 10 most abundant proteins–which include antibodies, the iron transporter called transferrin, and fibrinogen–account for more than 90 percent of all plasma protein. Many other proteins are present at extremely low relative concentrations. For example, albumin’s concentration is 5 to 10 billion times that of the immune system signal interleukin-6.
All broke up
Researchers are sorting out the best way to search for additional plasma proteins. Some add enzymes to their plasma samples to prevent the natural breakdown of proteins, yet others argue that these enzymes may hinder discovery. Investigators are also divided on whether to remove the albumin before they analyze a sample. That strategy might aid identification of low-abundance proteins, but it runs the risk of accidentally discarding plasma proteins that adhere to the abundant molecule. “Albumin is a sticky protein,” notes Pounds.
Therefore, in their recent analysis, he and his colleagues didn’t remove albumin from plasma samples. They did, however, seek to simplify their analysis by stripping the plasma of clotting factors. This creates a liquid commonly known as blood serum.
Also, by running the liquid over beads coated with a molecule that binds antibodies, they removed most of these immune proteins. They then added the enzyme trypsin to carve up the remaining proteins into small, but still recognizable fragments, or peptides.
The scientists next used a series of chromatography techniques that disperse peptides in a liquid or gas environment rather than on slabs of gel. This process separates albumin-derived peptides from those generated by other proteins, says Pounds.
Finally, the scientists injected each group of peptides into a tandem mass spectrometer, a device that weighs each peptide and then shatters it into its component amino acids and weighs each of them. From that information, the researchers deduced the amino acid sequence of each peptide and then identified the blood proteins that it came from.
When Pounds’ team applied this complex procedure to a few drops of plasma from a healthy woman, it identified 490 proteins, including some low-abundance ones such as human growth hormone and prostate-specific antigen. In their analysis, reported in the December 2002 Molecular and Cellular Proteomics, Pounds and his colleagues describe finding most, but not all, of the previously documented plasma proteins.
One protein that was missing was C-reactive protein, whose elevated presence in the blood may serve as a predictor of heart attacks (SN: 12/7/02, p. 364: Protein may signal heart problems). The plasma donor’s heart may have been so healthy that she had little C-reactive protein in her blood, or some facet of the group’s approach may have obscured the protein’s detection. For example, some peptides aren’t easily broken apart in a mass spectrometer.
Two proteins that typically function in the retina were unexpectedly found in the plasma. The woman might have some retinal damage that caused the proteins to leak into her bloodstream, speculates Pounds. Another explanation is that these proteins function in other tissues where scientists haven’t yet detected them.
Plasma portrait
Pounds says that his group’s work is just a first step toward compiling the master list of proteins in plasma. Members of the international Plasma Proteome Project are testing many other analytical strategies using chromatography and mass spectrometry, notes Omenn. “The whole idea is to have a global effort that characterizes the advantages and limitations of various technologies,” he says.
N. Leigh Anderson predicts that the number of known plasma proteins will double by the end of this year.
Once a low-abundance plasma protein is identified, it’s still a formidable challenge to develop an inexpensive and effective assay to detect it or measure its concentration. Such assays are needed if the protein is ever to be used to diagnose or predict a disease.
“Only a handful of proteins are currently used in routine clinical diagnosis, and the rate of introduction of new protein tests approved by the [Food and Drug Administration] has paradoxically declined over the last decade to less than one new protein diagnostic marker per year,” N. Leigh Anderson and Norman G. Anderson note in their review.
A promising advance on the horizon may speed development of such tests. Several companies are rushing to create simple devices that can detect hundreds if not thousands of proteins at a time. Similar in concept to the so-called DNA chips that were invented and quickly marketed about a decade ago (SN: 3/8/97, p. 144), these protein chips are wafers dotted with microscopic pools of molecules such as antibodies. As DNA chips enable scientists to monitor the activity of thousands of genes, protein chips could show the presence of thousands of proteins in a single blood test. When a liquid such as plasma is run over a protein chip, the antibodies bind their target protein, revealing its presence.
With such chips, suggests Pounds, researchers will devise better diagnostic assays for cancer. Whereas a single marker, such as prostate-specific antigen, may mislead a physician, simultaneous readings of multiple plasma proteins should be more definitive.
This wealth of new plasma-protein information could pose a challenge, however. “Physicians are not accustomed to looking at 10 different proteins in a certain pattern as an indication of disease,” notes N. Leigh Anderson.
Ultimately, through the analysis of proteins in plasma, researchers may capture a “molecular image” of a person’s normal physiology or of a disease state, says Pounds. They will also be able to study how a person’s diet, genetic background, lifestyle, and environment influence the contents of plasma, he adds.
“If we have all this information, we’ll probably be in a much better position to identify and design new drugs,” says Pounds. “We think there are thousands of proteins in plasma, and there’s a lot of opportunity here to improve human health.”
****************
If you have a comment on this article that you would like considered for publication in Science News, please send it to editors@sciencenews.org.
To subscribe to Science News (print), go to https://www.kable.com/pub/scnw/
subServices.asp.
To sign up for the free weekly e-LETTER from Science News, go to http://www.sciencenews.org/subscribe_form.asp.