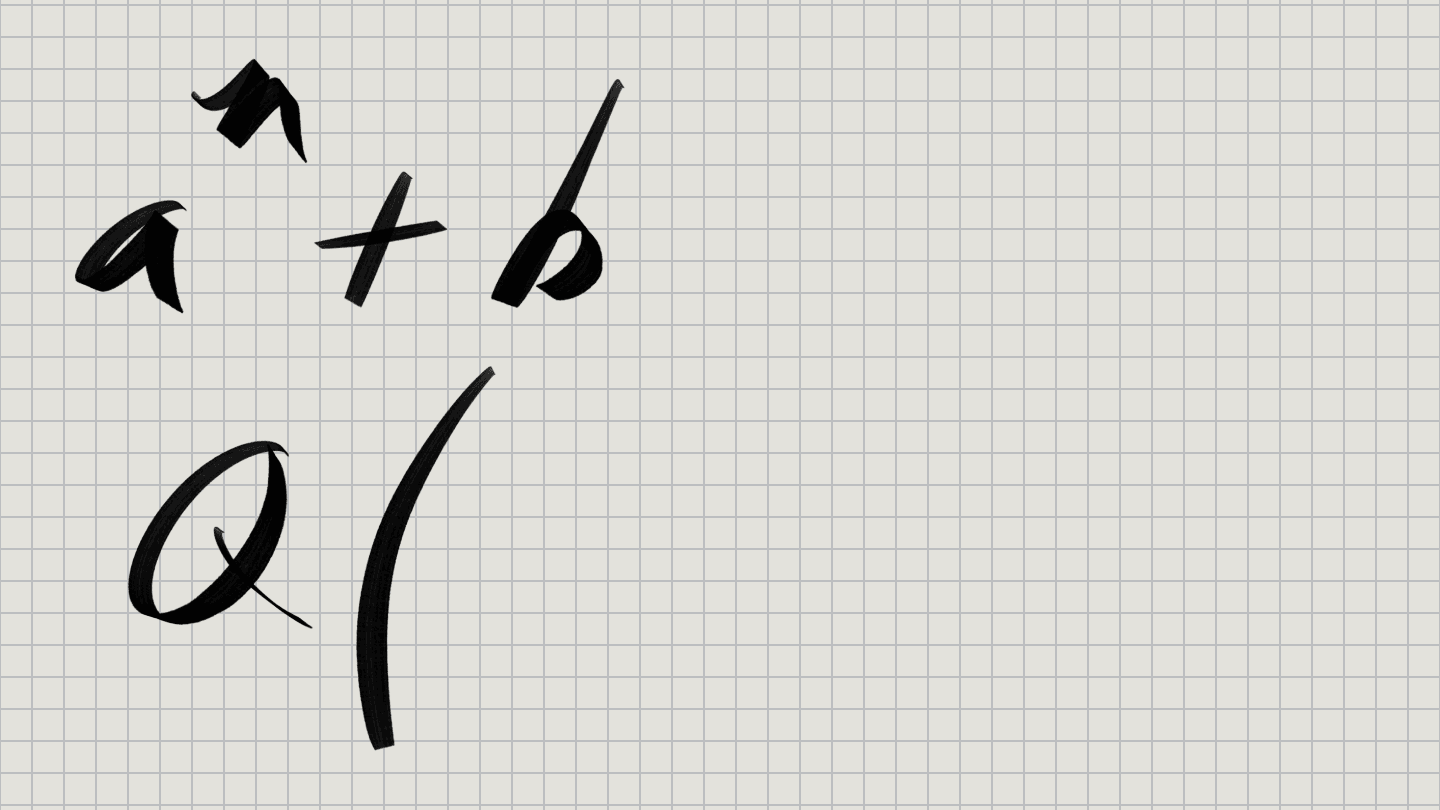It was all very nice when Watson and Crick figured out the structure of DNA more than 50 years ago, but since then, biologists have been trying to figure out how the genetic code works. They have a lot of data – this gene turns on that gene, these two genes turn off this other one. But together, all those nice data turn into a great snarl, a spaghetti-like mess of relationships between one gene and another. What does it all mean?

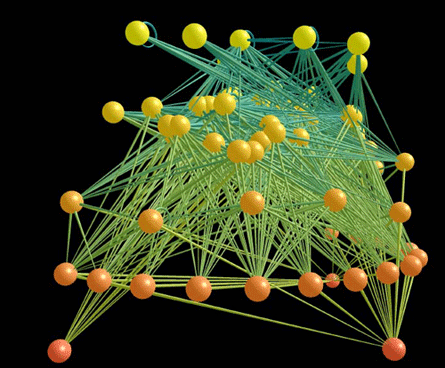
A radical new approach to modeling networks may help molecular biologists sort their spaghetti. At the same time, it may help computer scientists understand the organization of the Internet, ecologists untangle food webs and CIA agents ferret out terrorists. Furthermore, the technique can even predict new connections scientists hadn’t known about in each of these fields.
“What we want is the operating manual for the cell or organism,” says Aaron Clauset of the Santa Fe Institute in New Mexico. “To get that, we need to explain the structure at a variety of levels.” One group of genes, for example, might work together to form the retina while a larger group functions to form the eye and a still larger group constructs the entire nervous system.
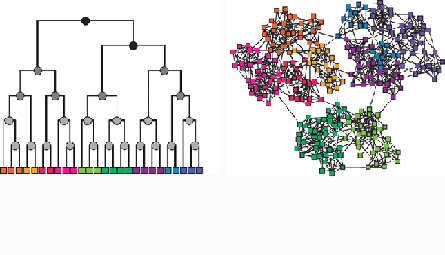
So Clauset, together with Cristopher Moore of the University of New Mexico and physicist Mark Newman of the University of Michigan, created a class of models designed to expose this kind of structure within networks. They started with a simple family tree where each branch of the tree defines a community, with sub-branches forming subcommunities. To allow for connections between communities, at each branch point they assigned a probability that a node on the right side of the branch would be connected to a node on the left.
This approach of assigning different probabilities allowed for many different types of communities. Often, networks have a clumpy structure, with a lot of connections inside the clumps and fewer between them. Take social networks, for example: If you and I are both friends with Anne, we’re probably friends with one another. That makes social groups pretty tightly integrated. At the same time, social groups tend to be distinct: the cool kids rarely mix with the nerds.
This kind of clumpy structure emerges naturally from the researchers’ class of models when the probabilities of a connection are greater between nodes that are closer on the tree than more distant nodes. If you and I are both friends with Anne, she probably lies close to both of us on the family tree. That means we’ll be close to one another as well, and we’ll likely have a connection of our own. The cool kids and the nerds, on the other hand, will be on different sides of the tree entirely.
Although this structure is common in networks, it’s not universal. Food webs, for example, don’t form clumps in that way. That’s because if you and I eat the same thing, we’re not more likely to eat one another – in fact, we’re probably less likely to do that. Still, predators of the same creature form a kind of community. Clauset’s class of models can capture this too, by making the probabilities of a connection between closely related nodes less than those at more distantly related ones.
The challenge lies in identifying the right structure to use for a particular dataset – choosing the right shape for the tree and the right probabilities. “The neat thing is that we don’t decide upfront which kind of clumping we expect for a particular network,” Clauset says. “We let the data tell us instead.”
To do that, they used the magic of machine learning. Given a way of determining which model is better than another, computers can use fancy search techniques to find the best ones, even when there are way too many models to test them all. The computer then averages together the best models, getting a combined model that is better than any single one.
Clauset and his colleagues set high standards for their models. They didn’t just want models that reproduce the original data faithfully. They also wanted ones that are robust, so that new data won’t be likely to change the model radically.
So to decide which models were best, the researchers played a little trick: They hid some of the links when they built their candidate models. Then they pulled out those hidden links. If the model was better than chance at predicting what was hidden, they knew it had learned something about the structure of the network.
This testing method has a remarkable side effect. “It can be used as an interactive oracle,” Clauset says. Usually, researchers don’t know all the links in the network they’re studying. Food-web biologists, for example, know some of the things each animal eats, but not all. Clauset’s model can predict missing links just from the topology of the network, suggesting the odds are good that, say, an animal eats a particular plant even though the biologists have never observed that. The biologists can then use those predictions to guide their fieldwork.
The researchers tested their method on three well-studied networks: the food web of a grassland ecosystem, the metabolic interactions of the bacteria that causes syphilis and the associations between the terrorists that carried out the September 11 attacks. Clauset and his colleagues found that their method was able to identify the subcommunities in each of these networks with great accuracy just from the structure of the network. They published their results May 1 in Nature.