Physicists join immune fight
Principles beyond biology may help explain how the body battles infection
A runny nose, sore throat and fever may send you running to your doctor — a perfectly satisfactory strategy if all you are seeking is relief. Applying their knowledge of biology, doctors can analyze your symptoms and then prescribe the best drugs to alleviate them. But if you want to know what’s really going on inside your body, consider asking a physicist.
Through the years, biologists have identified and described the various cells that orchestrate the immune system’s response to infection, leading to effective vaccines and treatments for many diseases. Mathematical experts have helped to get a handle on the numbers, revealing how many cellular players there are and where they go, as well as showing how quickly the body clears an infection or how long it takes to acquire resistance to a drug. But these approaches haven’t yet revealed the full playbook directing the coordinated activities of all involved.
Fighting an infection requires scores of players, each with a job to do. Some cells patrol the body, seeking out and devouring invaders. Others send signals, receive messages or destroy infected or mutated body cells. As the various components in the immune system carry out their work, they interact over multiple levels and time periods, says Arup Chakraborty, a chemical engineer at MIT.
Such complexities make it difficult to intuit the underlying mechanistic principles involved through experiments or simple calculations alone, he says. So now physicists are being called into the game, applying laws governing movement and matter to explain the complex goings-on. Using approaches from statistical physics — a field that employs probabilities to predict and describe the outcomes of events that arise from the interactions of many linked variables — scientists are developing new models to study the play-by-play maneuvers that occur during a full-blown immune response.
By coupling laboratory experiments with this new approach, researchers are beginning to uncover some of the key events in immune signaling. Chakraborty and his team have studied how some immune cells operate in an on-or-off digital manner and have explored how the cells learn to distinguish invaders from the body’s own tissues. Current studies using the statistical physics approach are looking at how immune cells create and keep memories of past infections. Others are developing new theoretical models to explain how immunity adapts to keep up with ever-changing pathogens.
Understanding how immunity works as a single system may help scientists find new ways to manipulate and control it, leading to more effective vaccines or improved treatments for persistent autoimmune disorders.
By the numbers
The immune system’s first line of defense is to prevent bacteria, parasites, viruses and other infectious agents from entering the body. Skin and the cells that line the nose and stomach serve as a front line, keeping out and wafting away dirt, dust and germs. Specialized cells also patrol the bloodstream, attacking and devouring any germs that get through.
If a bug manages to outwit this system and infect a cell, the adaptive immune system kicks in. This special team serves as a backup defense, designed to identify offenders and produce legions of cells ready for combat. The process starts when white blood cells swoop in to capture and kill the invaders, taking their remains to the lymph nodes. There, protein fragments of the invading pathogen are displayed on the surface of antigen-presenting cells. Serving as a red flag, the fragments identify the bug and signal the immune system to attack cells infected by that agent. Leading this attack are the T cells, white blood cells that work to neutralize infectious agents.
Generally, the various immune cell types exist in small numbers. But under threat, cell numbers can quickly expand into the millions. These cells use chemicals to talk and coordinate an attack.
“It’s a game of numbers as much as it is a game of molecular and cellular biology,” says biologist Rustom Antia of Emory University in Atlanta.
Alan Perelson, a theoretical immunologist at the Los Alamos National Laboratory in New Mexico, was one of the first to show how mathematical models could be used to study immune changes occurring during infection. In the early 1990s, his team developed models to track how the number of T cells shifts in response to HIV, the virus that causes AIDS.
At the time, many scientists considered AIDS to be a slow-acting viral infection because it takes many years — on average a decade — for symptoms to appear.
Using models to analyze the amount of virus in the body, Perelson’s group showed that the virus is, in fact, quite active during this seemingly latent period. The findings revealed that once a person is infected, the virus is cleared from the body quickly. But, in the absence of drug therapy, it multiplies just rapidly enough to keep up with this clearance.
Since Perelson’s work, advances in instrumentation have made it possible to collect much more quantitative data. “I think people are starting to recognize the value of having quantitative information,” Perelson says.
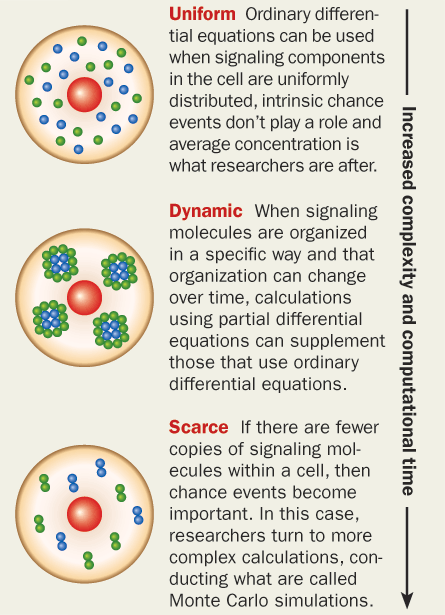
While simple mathematical models that rely on a limited amount of data can track the movements and actions of large numbers of players, explaining the play-by-play actions of smaller teams within the team is much harder. In the same way that football has special teams that are called in only during a punt or a field goal attempt, the immune system has special units — made up of molecular players — that get called in for certain situations.
Chakraborty says such orchestration creates a “hierarchy” of organized cooperative processes. With so many interacting components, calculating the behavior of any particular individual is not straightforward.
This is where statistical physics can help. By using equations that take all the interactions into account, researchers can estimate the likelihood that a certain result will occur.
Sending out an SOS
Chakraborty, a chemical engineer and biochemist by training, became interested in immunology after hearing about a paper on the immunological synapse, a crucial communications point for T cells and antigen-presenting cells. Studies had shown that when the two cells meet, various receptors and molecules bind across the synapse, organizing into a doughnut-shaped pattern. But researchers couldn’t figure out what function the structure served.
After thinking about its possible signaling function, Chakraborty developed a series of computer simulations to look at all the possible biochemical events that the synapse might influence. The simulations showed that the synapse acts as an adaptive control device, ramping up to enhance T cells’ sensitivity to an antigen, but backing off so as not to overstimulate or kill the T cell.
Once in the immunology game, Chakraborty got hooked. He is now bringing his engineering background to another question: How do T cells determine their course of action?
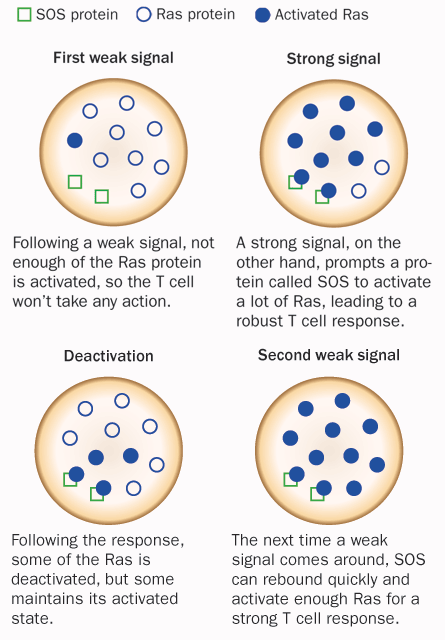
T cells aren’t fickle. As they go about their activities and respond to the environment, the cells make friend-or-foe distinctions on the fly. A timid T cell could invite all kinds of germs and misery, while an overzealous one could cause allergic reactions and harm the body’s own tissues. The deliberations start when T cells and antigen-presenting cells meet in the lymph nodes, causing a series of molecular events leading to the activation of a protein called Ras.
Arthur Weiss, an immunologist at the University of California, San Francisco, and his colleagues had done a lot of work identifying the pieces of the signaling machinery, but the intricate regulation of the T cells’ decisions to mobilize was a puzzle. T cells contain two types of Ras activators: a molecule called SOS and another called Ras guanyl nucleotide-releasing protein, or RasGRP. The scientists hypothesized that SOS worked to amplify T cell receptor signaling while RasGRP played a priming role for SOS.
When Weiss described the complex signaling system at a scientific meeting in 2006, Chakraborty was in the audience. He approached Weiss after the presentation to suggest a way the system might work, based on a physical phenomenon called hysteresis. In hysteresis, a system’s response is determined by the last action exerted on it. Iron, for example, remains magnetized for a brief period of time even after the magnet is removed — and responds according to that magnetization.
Chakraborty proposed that the chain of events that led a T cell to go forth and conquer involved a type of hysteresis; the response to signals from antigen-presenting cells was based on recent encounters with other antigen-presenting cells. His group then developed simulations to mimic this type of interaction. The simulations showed that the interplay of Ras and SOS creates a positive feedback loop — a condition in which a little bit of Ras activation by SOS makes a future, more robust activation more probable.
In further experimental studies, Chakraborty, Weiss, Jeroen Roose of UCSF and colleagues showed how the second Ras activator, RasGRP, can prime SOS, amplifying its signal. Through this mechanism, the T cells not only exhibit hysteresis, but also work in an on-or-off fashion, much like a digital circuit in a computer, the team reported in 2009 in Cell.
Weiss says Chakraborty’s insights and understanding of physical systems allowed the team to extend its understanding of the immune signaling system.
“He was able to take experimental observations that largely already existed and provide a computational model that could explain how different quantitative parameters in signaling could give qualitatively different outcomes,” Weiss says.
The model also showed how small tweaks in the signaling process could give rise to large effects, Weiss says. “That’s something that I think we biologists don’t understand because it’s not the kind of training that we generally have.”
Dealing with frustration
Recently, Chakraborty and his team developed another type of model to study the education process that T cells go through when learning to tell the body’s cells from those of invaders. The model, first described in 2009 in Physical Review Letters and later in May 2010 in Proceedings of the National Academy of Sciences, is based on an idea from statistical physics called “frustration.”
A frustrated system in physics — much like any of life’s trying circumstances — occurs when a system has to make many players happy simultaneously, but has no power to change itself or its conditions in order to make that happen.
Such is the case when T cells learn to distinguish between fragments of foreign proteins and bits of the body’s own proteins, or self-peptides. T cells that bind too tightly to self-peptides die off. Those that bind weakly come out as winners, and move out into the bloodstream to perform their killer duties.
The frustration enters because the T cell’s receptor can’t change its configuration, yet it has to learn to bind weakly to a variety of different self-peptides. “It’s sort of like you have to make friends with diverse people who are very different,” Chakraborty says. “It’s very hard. It takes a certain personality to do that.”
Last May in Nature, Chakraborty and colleagues described how this learning process could be a contributing factor to how people with certain genes can control HIV better.
While learning to make friends, T cells also remember past enemies. Emory’s Antia is working with Rafi Ahmed, head of Emory’s vaccine center, to model ways in which long-term memories of opponents are stored so they can be recognized the second time around. Antia describes the model as a bucket filled with memory cells against different pathogens.
“When the immune system encounters a new pathogen, there are one of two things it might do,” he explains. “One, it can expand the size of the bucket, or two, it can throw away things that were in the bucket already to make room for the new memories.”
Because people are a finite size, researchers first theorized that the bucket was limited in size as well. Indeed, models showed that once the bucket is filled, the immune system begins tossing some of the preexisting cells. Rather than tossing the earliest memories first, the models suggest that the rule for discarding is to reach into the bucket and choose cells at random.
The scientists then developed a model to see if the bucket could grow. The simulations called for adding huge numbers of new immune memory cells to see how the preexisting memory would decline.
“We actually thought the test would be trivial,” Antia says. “We thought it would just test our previous model.”
But the second model, published in Nature in early 2009, showed that the immune system’s repertoire has the ability to expand. Antia is now working with experimental biologists to find ways to test the predictions in animals.
Back to the body
While statistical physics can inform basic biology, it also has something to say about preventing disease.
Biomathematician Michael Deem of Rice University in Houston became interested in studying the immune system 15 years ago when he rolled up his sleeve to get his annual flu shot. While administering the shot, the nurse told Deem that getting a flu shot in any given year could, in certain circumstances, make you more susceptible to the flu the following year. Viruses such as the flu are constantly evolving into new variations. A minor change in a strain that the body has learned to recognize can allow the new bug to slip right past.
The incident pushed Deem to develop theoretical models that could more accurately predict which vaccines work best against ever-evolving flu strains. In 2006 his group introduced a model to look at variations in amino acids in the regions of a virus that the immune system recognizes, called epitopes. The model is based on random energy models from physics that are used to study complex systems with many “metastable” points, or states in which the system can get stuck.
Using information from the virus, vaccine and antibody structures, scientists can predict tendencies for amino acid evolution in the epitopes, and develop new vaccines accordingly. Deem says his new model, detailed in February 2010 in the Annual Review of Chemical and Biomolecular Engineering, may better predict the effectiveness of a vaccine than do the animal models currently used.
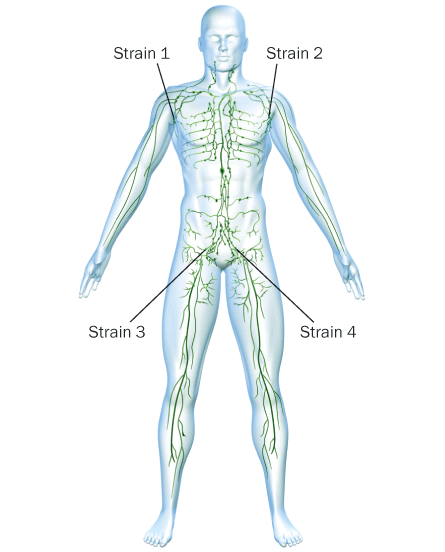
Deem is also using his model to find ways to counter diseases caused by viruses that have multiple strains. Dengue fever, for example, can be caused by any one of four strains of the virus. People who are infected with one strain are more likely to get dengue hemorrhagic fever if infected by one of the other three strains. Vaccinating for all four strains in a single vaccine doesn’t work, though, because the immune system will focus on only one or two dominant strains.
While simulating how such dominance occurs, Deem’s team came up with an idea: Because the immune system does its looking in lymph nodes located throughout the body, why not inject different variations of the virus into different areas of the body? The process would eliminate the competition for the immune system’s attention because each of the four different strains would go to a different lymph node, Deem says.
Recent experiments in animals suggest that the multivaccination approach works. In one experiment, monkeys were immunized with four different strains of dengue virus in four different injections, one in each leg and each arm.
For a short period of time, a few days or so, the antibodies and T cells produced in response to each strain remain localized at a lymph node near the injection site. Eventually, though, the T cells from each lymph node circulate throughout the body, giving protection to all four strains.
If Deem has his way, future vaccinations may take advantage of such principles, nudging physics a little further into the doctor’s office.