Quite a Switch
Bacteria and perhaps other life forms use RNA as environmental sensors
Let’s start at the very beginning, several billion years ago, when the first specks of life began to crawl across the barren planet that’s now called Earth. There were no cameras to record images of these early life forms, and it’s unlikely they left any physical fossils, so it’s all but impossible to know much about the primitive organisms. That hasn’t stopped scientists from speculating, however. One of the most provocative notions about Earth’s first life is that it was far different from that today.
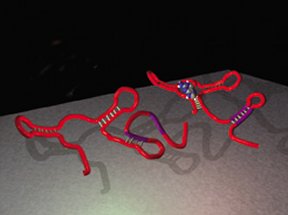
Modern life has settled on a tried-and-true plan, with the starring molecules consisting of proteins and deoxyribonucleic acid (DNA). Cells now depend upon genes made of DNA to store the blueprints for proteins, which have many duties: enzymes, signaling molecules, environmental sensors, and regulators of gene activity, for example. But some scientists argue that there was a time before proteins and DNA. This lost world was ruled by ribonucleic acid (RNA), the single-stranded relative of DNA.
Instead of proteins and DNA, the ancient organisms would have relied on strands of RNA to perform chemical reactions or to pass information from one generation to the next. If such an RNA world did once exist, the chemical must be much more versatile than most scientists have deemed possible.
“If you’re going to be a complex organism in the RNA world, you’re going to want to be able to click things on and off. You’re going to want to sense the environment and respond to it. All of these things would have had to be done by RNA because there were no proteins,” says Ronald Breaker of Yale University.
He and several other scientists may now be unearthing remnants of this ancient RNA world within the genes of modern microbes. In a series of recent publications, they’ve shown that strands of RNA can by themselves act as environmental sensors that regulate gene activity.
While studying bacteria, these researchers have identified RNA segments that detect shifts in temperature (see “Some Like It Hot,” below) or monitor concentrations of vitamins, amino acids, or other small molecules within a cell. In response to changes in the environment, these RNAs alter their shapes, thereby stopping or starting production of the proteins they encode.
The discovery of these riboswitches, a name coined by Breaker’s research team, presents scientists with a new way to explain how bacteria—and probably more-sophisticated organisms—regulate their genes. Investigators are already exploring ways to exploit riboswitches. Some envision incorporating riboswitches into genes being used for gene therapy. Others see bacterial riboswitches as potential targets for new antibiotics.
“In just the last 2 years, at least seven families of riboswitches have been uncovered. I think that’s just the tip of the iceberg,” says Evgeny Nudler of New York University Medical Center. “There could be many more. They’re being uncovered with increasing frequency.”
You can’t beat nature
Biologists know RNA best as a simple courier. For a modern cell to produce a protein, it typically uses the information encoded by a gene’s DNA to make a strand of RNA. Known as messenger RNA (mRNA), this strand carries the blueprint for a gene’s protein to factories within a cell called ribosomes.
Researchers have gradually recognized other roles for RNA, fueling suspicions about the ancient RNA world. For example, organisms produce RNAs that can act like protein-based enzymes. Scientists, seeking to improve on nature, have even synthesized these so-called ribozymes in test tubes (SN: 8/7/93, p. 90; 4/7/01, p. 212: Available to subscribers at RNA world gets support as prelife scenario).
A biochemist, Breaker has worked on synthetic ribozymes for many years. Recently, he’s explored what else artificial RNA can do. In particular, he and other investigators have begun to create strands of RNA that can target and bind to specific small molecules, much as the proteins known an antibodies do.
These RNAs, dubbed aptamers, could prove useful in many ways, including diagnostic tests similar to those that now use antibodies. To find aptamers, scientists cast a target molecule into solutions containing millions of subtly different synthetic RNAs and examined the ones that stuck to the target.
But has nature already invented aptamers? In other words, do cells employ RNAs to bind to small molecules? Some biologists over that past decade have proposed that natural RNAs might regulate gene activity by binding to metabolites, the small molecular products of a cell’s metabolism. In one scenario, the mRNAs for enzymes used to make a vitamin would sense that a cell had enough of the vitamin. Those mRNAs might then somehow prevent themselves from being translated into the enzymes.
The success at synthesizing artificial aptamers hinted at the existence of natural aptamers. “It was a ripe idea that made a lot of sense,” says RNA researcher Gary Stormo of Washington University Medical School in St. Louis.
Because there were no definitive examples in the scientific literature of natural aptamers, Breaker looked for instances in which microbiologists were at a loss to explain how a metabolite controlled a gene’s activity.
He found a handful of cases in which researchers had established that a metabolite turned off production of a protein and that a portion of the gene’s mRNA was central to that control. Most scientists, however, had assumed that a protein sensed the metabolite and then bound to the mRNA.
“The dogma in the field was that [gene] regulation is done by proteins,” says Stormo.
In contrast, Breaker’s team suspected that the mRNAs themselves were sensing the metabolites. “We believed biologists had seen riboswitches in action without knowing it,” Breaker says.
To back up this suspicion, Breaker and his colleagues initially focused on a gene in the bacterium Escherichia coli. The gene, which encodes a protein that transports vitamin B12 within the microbe, appears to stop making the transporter when a lot of the vitamin is available.
In 2002, Breaker’s team established that the mRNA for the transport protein acts as a riboswitch. It latches directly onto vitamin B12 without the aid of any protein, and this binding alters the mRNA’s structure so that it can’t direct production of the transport protein. As a result, even though the gene for the transport protein is active, sending out its mRNA, the protein won’t be produced when the microbe has plenty of vitamin B12.
In a string of papers that quickly followed, Breaker’s team unveiled riboswitches in E. coli and other bacteria that respond to different vitamins and metabolites. Around the same time, Nudler’s group published data documenting some of the same riboswitches. He had been inspired to look for them by colleagues in Russia who had searched in vain for metabolite-binding proteins that control certain bacterial genes.
“We were scooped,” Nudler ruefully admits.
Both groups, for example, identified a natural riboswitch that recognizes flavin mononucleotide (FMN), a derivative of vitamin B2. When abundant FMN is present, the switch shuts down production of several enzymes that synthesize vitamin B2.
As it turns out, Breaker’s team had previously synthesized an aptamer that binds to this metabolite. The natural riboswitch is five times as large as the synthetic one, looks completely different, and does a much better job at picking out FMN from other molecules, Breaker says.
Off and on
Most riboswitches discovered so far act as off switches. When something binds to the mRNA, the strand’s structure changes in a way that prevents the cell from using the RNA to manufacture its protein. The prevalence of such switches makes sense, according to Breaker. “Biology has the greatest need for shutting things off in response to rising metabolite concentrations. If you’ve made enough compound X, you want to shut down the genes that make compound X,” he explains.
In studies of the bacterium Bacillus subtilis, however, Breaker and his colleagues recently uncovered the first riboswitch that, when triggered, promotes a protein’s production. This on switch responds to adenine, a component of RNA, DNA, and some other molecules. The switch is in an mRNA that directs production of a protein that sits in cell membranes and pumps adenine out of the cell. When adenine binds to the riboswitch, the mRNA changes into a shape that enables the cell to make the pump, the researchers reported in the January Nature Structural and Molecular Biology.
“It’s just a variation on a theme,” says Breaker. “It’s one of the rare instances in which you want to increase the expression of a gene in response to rising concentrations of a metabolite.”
The most recent riboswitch unveiled by Breaker’s team, described in the March 18 Nature, ups the ante in terms of complexity. The switch works in concert with a ribozyme contained in the same mRNA molecule. That molecule encodes an enzyme that produces the sugar glucosamine-6-phosphate. In response to the sugar, the switch triggers the ribozyme part of the mRNA, which then cleaves the mRNA in such a manner that the sugar-producing enzyme can’t be made.
Riboswitches have been identified in many bacteria, but Japanese scientists just identified the first riboswitch in a fungus. This finding is important because fungi are eukaryotes, the group of organisms that includes plants, animals, and people. Indeed, researchers scanning plant and fungi genes have found DNA sequences that they predict could produce mRNAs with riboswitches.
“Once the first riboswitch is found in a higher eukaryotic organism, that will trigger an avalanche of research,” says Nudler.
It’s easy to see that denizens in the ancient RNA world needed riboswitches, but why would eukaryotes and bacteria today continue to use RNA, rather than proteins, to control gene activity? Having mRNAs with built-in switches may offer cells a way to avoid wasting time and energy on production of regulatory proteins. “It’s expensive [for a cell] to synthesize a new protein,” notes Nudler.
The riboswitch “seems a very efficient way for microorganisms, in particular, to regulate gene expression. It’s every bit as effective [as proteins] and saves you a few steps,” agrees Robert Weisberg of the National Institutes of Health in Bethesda, Md., who studies RNA’s role in gene regulation.
In fact, given that riboswitches seem to work so well, Weisberg now wonders why bacteria began using gene-regulating proteins in many cases.
The lost world
Biologists are eager to put riboswitches to work. Breaker is working with Archemix, a company in Cambridge, Mass., that seeks to use combination riboswitch-ribozymes in sensors and diagnostic tests. A manufactured riboswitch might someday detect minuscule amounts of the anthrax bacterium, for example.
Riboswitches could also improve physicians’ control over gene therapy, a still-experimental strategy in which genes transferred into a person’s cells cure a disease. Breaker envisions modifying these transferred genes to incorporate a riboswitch into their mRNA. The switch would permit the production of the gene’s protein only in the presence of a certain drug. A patient could thus activate the gene by taking a pill or suspend therapy by refraining from the drug.
Antibiotics that target riboswitches are also under discussion. Since these RNA segments control production of essential metabolites in bacteria, scientists could develop compounds that mimic the triggers of the switches. Such a compound would trick the germ into shutting down the synthesis of a needed metabolite. The bacteria would, in essence, starve itself to death.
Scientists may have already found a chemical that does this. In the Nov. 1, 2003 Genes and Development, Breaker and his colleagues described a riboswitch that binds to the amino acid lysine. They reported that a previously identified bacteria-killing compound called S-(2-aminoethyl)-L-cysteine appears to work by flipping this switch and fooling B. subtilis into thinking it has enough lysine.
Whether or not physicians can exploit riboswitches, Breaker finds it amazing that these RNA sensors persist in modern cells.
“Riboswitches are like a little chunk of the lost RNA world,” he says. “It makes you wonder how many other chunks of the RNA world are not entirely lost.”
Some Like It Hot
Bacterial RNA senses body heat, turns on virulence genes
Like many other bacteria, the foodborne Listeria monocytogenes knows when it’s found a warm home. When the microbe, which can cause severe illness in pregnant women and people with weakened immune systems, is inside a human body, it kicks into high gear its genes for multiplying. It also revs up these genes when grown at body temperature, 37°C, in a laboratory dish. The genes fall silent at 30°C.
Some bacteria use proteins, or even DNA strands, as thermosensors, but L. monocytogenes depends upon a different strategy. The microbe uses RNA to sense when it’s inside a host, a research group headed by Pascale Cossart of the Pasteur Institute in Paris has found.
The investigators knew that L. monocytogenes relies on a DNA-binding protein called PrfA to turn on its virulence genes. Not surprisingly then, bacteria grown at 30°C contain much less PrfA than do ones kept at 37°C in the lab.
The explanation for this difference lies in the messenger RNA (mRNA) that carries the code for PrfA from L. monocytogenes‘ DNA to the bacterium’s protein-building ribosomes. Most of the mRNA strand holds the information needed to make the protein, but there’s an extra portion that turns out to be thermosensitive. At 30°C, this part folds up into a structure that prevents the mRNA from binding to a ribosome. As a result, the bacterium makes little or no PrfA.
At 37°C, however, the thermosensitive bit of RNA assumes a shape that permits the mRNA to interact with the ribosome. Voilà, the microbe can manufacture PrfA.
Cossart and her colleagues, who reported their conclusions in 2002, showed that they could make subtle alterations in the critical area of the mRNA and change the temperature at which L. monocytogenes turns on its virulence genes. For example, they created a strain of the bacterium that is virulent at 30°C.
The investigators even transferred the original thermosensor into another microbe. They modified a gene for green-fluorescent protein so that its mRNA incorporates the sensor. When the new gene was added to Escherichia coli, the altered bacteria glow green at human-body temperature but not under cooler circumstances.
“I would be very shocked if there aren’t many, many ways in biology that RNA motifs are used for thermosensing,” says Ronald Breaker of Yale University.
In a 2002 commentary on Cossart’s work, noted John C. Newman and Alan Weiner of the University of Washington in Seattle, suggest that this way of sensing heat represents a vulnerability for L. monocytogenes and perhaps other disease-causing bacteria. They speculate that small molecules that bind to PrfA’s mRNA and stabilize it in its low-temperature, unproductive form could stop a germ’s virulence, thereby acting as a novel class of antibiotics.