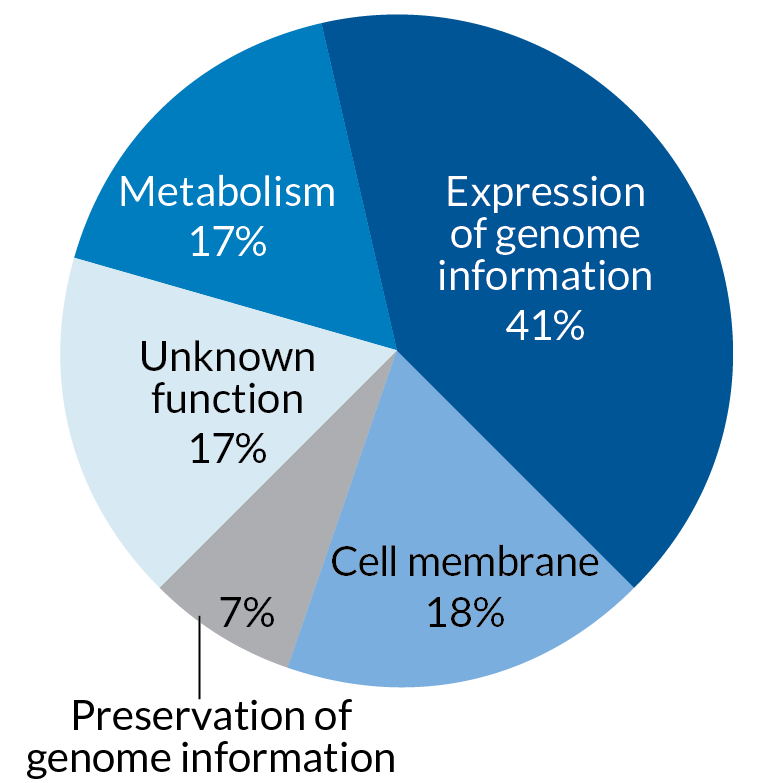
STRIPPED DOWN A newly-created bacterium dubbed syn3.0 has the smallest genome needed for free living. It contains just 473 genes. Of those, 149 have unknown functions.
Credit: E. Otwell; Source: Hutchison et al/Science 2016.
Scientists have built a bacterium that contains the minimal genetic ingredients needed for free living.
This bacterium’s entire set of genetic blueprints, its genome, consists of only 473 genes, including 149 whose precise biological function is unknown, researchers report in the March 25 Science.
The newly-created bacterium contains a minimalist version of the genome of Mycoplasma mycoides. Mycoplasma already have some of the smallest known genomes. M. mycoides used in the experiments started with 901 genes. In comparison, other bacteria, including E. coli, may have 4,000 to 5,000 genes. Humans have more than 22,000 genes, although not all are necessary (SN: 4/2/16, p. 18).
In 2010, researchers at the J. Craig Venter Institute in La Jolla, Calif., replicated the entire genome of M. mycoides and popped it into a cell of a different species, Mycoplasma capricolum, creating what some people called the first synthetic organism (SN: 6/19/10, p. 5). The new work strips the M. mycoides genome down to its essential elements before transplanting it to the M. capricolum shell, producing a minimal bacteria dubbed syn3.0.
Researchers hope syn3.0’s uncluttered genome will teach them more about the basics of biology. Such minimal genome bacteria also may be chassis on which to build custom-made microbes for producing drugs or chemicals.
J. Craig Venter, founder of the nonprofit institute, and a team of researchers there led by Clyde Hutchison III and Daniel Gibson initially set out to design an organism based on a core set of about 300 genes that researchers surmised a microbe would need to survive on its own. But when the researchers tried to bring their computer creations to life, “every one of our designs failed,” Venter said in teleconference with reporters. The failure was due to leaving genes of unknown function out of the mix. About 32 percent of the genetic ingredients ultimately needed to cook up even a simple organism were left out of the initial recipe because the researchers didn’t know what the genes did and didn’t understand their importance. Once those genes were mixed back into the batter, the bacteria sprung to life.
“I think we’re showing how complex life is in even the simplest of organisms,” Venter said. “These findings are very humbling” because they show that researchers still don’t fully understand even the minimal requirements for life.
That lack of knowledge is “frustrating after so many years of molecular biology,” says synthetic biologist Pamela Silver of Harvard Medical School. But the pared-down microbe may be a good platform for discovering what genes of unknown function do, she says.
Other researchers have attempted to make minimal genomes by stripping away one gene at a time. But the Venter group built their lean microbe from the ground up, synthesizing pieces of DNA that would later be stitched into a complete genome.
Drew Endy, a synthetic biologist at Stanford University, is among several scientists applauding the made-from-scratch approach. “Only when you try to build something do you find out what’s truly required. Too often in biology we end up with only data, a computer model, or a just-so story. When you actually try to build something, you can’t hide from your ignorance,” Endy said in an e-mail. “What you build either works or it doesn’t.”
At first, the bare-bones genome didn’t work. Some genes that appeared to be nonessential for life are really requirements, the researchers discovered. Those genes tended to have redundant functions with another gene. Researchers could remove one of those genes, but not both at the same time, just as knocking out one engine on a twin-engine jet will keep the plane airborne, but disabling both engines will lead to a crash, says Gibson.
Although syn3.0’s genome is far smaller than those of other free-living bacteria, it may not be the minimal genome for every independent organism in every situation. (Symbiotic bacteria living inside host cells may have fewer genes than syn3.0 does, but cannot survive on their own.) Other researchers have theorized that a minimal cell could consist of one single RNA-replicating gene inside a membrane, says geneticist George Church of Harvard University.
Starting with another organism or growing the bacteria under different conditions would probably lead to a microbe with a different minimal set of genes, says Jay Keasling, a synthetic biologist at the University of California, Berkeley. “The minimal genome is in the eye of the beholder,” he says.
Gibson and Venter agree that they have created a minimal genome, but not necessarily the minimal genome. Syn3.0 is streamlined, but still contains a few frills. The team kept several “quasi-essential” genes that aren’t strictly necessary for life, but allow the bacteria to grow fast enough to make them useful in the lab.