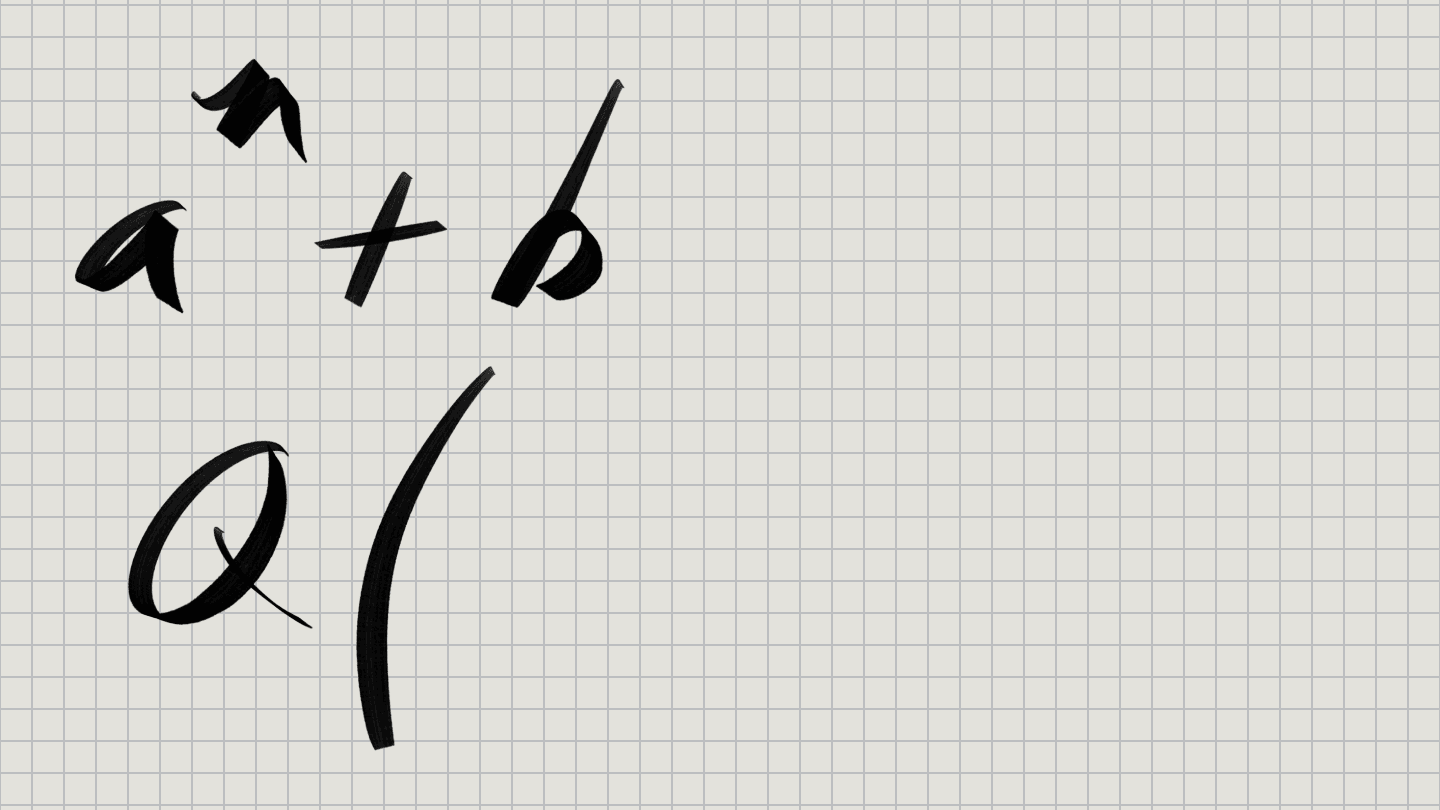It’s amazing how easily a loosely coiled rope can acquire a knot. The same sort of tangling can happen to a long hose lying in an untidy heap in a garden shed, earphone cords connected to a music player, and a necklace in a jewelry box.

Given that it normally takes some effort to create a knot, the seemingly spontaneous formation of knots in ropes and strings can appear puzzling.
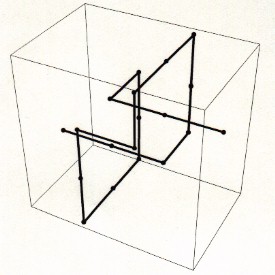
The secret of spontaneous knotting lies in the mathematics of self-avoiding random walks. One way to model such a walk in three dimensions is to track a walker standing at a vertex of a three-dimensional grid. The walker can take a step from one vertex to the next in any one of six directions. Randomly choosing each direction with equal probability, the walker traces a path that wanders from vertex to vertex, sometimes crossing itself and sometimes retracing steps.
In 1988, mathematician De Witt L. Summers and chemist Stuart G. Whittington proved that nearly all sufficiently long self-avoiding random walks on a simple cubic lattice contain a knotted pattern.
Such a mathematical model could apply to the long molecular chain of a polymer immersed in a solvent. Indeed, Sumners and Whittington were originally inspired to tackle the problem in order to better understand the tangling of polymer chains and the occurrence of knots in ring polymers.
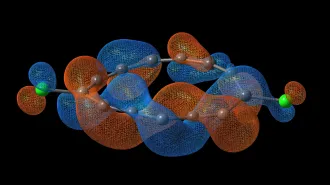
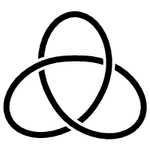
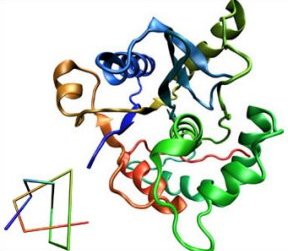
A protein is typically a long chain of amino acids, intricately folded into a compact package. Interestingly, although abundant and complex in polymers, knots are rare and simple in proteins. For the most part, knotted proteins contain trefoil knots (represented in diagrams with three crossings). Only three proteins have been found with knots that have four crossings (figure-8 knot).
Now, Peter Virnau, Leonid A. Mirny, and Mehran Kardar of the Massachusetts Institute of Technology have uncovered the most complicated knot yet discovered in a protein—one with five crossings. They describe their findings in the September PloS Computational Biology.
Only two distinct knots have five crossings.
The mystery is why a protein would have such a complicated knot and, more generally, why any proteins would have knots.
Of the 36 protein knots studied by Virnau and his coworkers, all are in enzymes. The five-crossing knot is contained in a protein called human ubiquitin hydrolase. This protein rescues proteins otherwise marked for destruction.
Normally, a protein destined for destruction within a cell is labeled with another protein called ubiquitin. Marked proteins are then shuttled to a cell structure called a proteasome, which takes in the protein and chops it up. Ubiquitin hydrolase, however, intercedes in this process and removes the offending ubiquitin, saving the comdemned protein.
A proteasome starts its job of degrading a protein by unfolding it—a task accomplished by threading the protein through a narrow pore. “Such threading unto the degradation chamber depends on how easily a protein unfolds, with more stable proteins being released back into solution and unstable ones being degraded,” Virnau, Mirny, and Kardar say.
The rescue protein’s own entangled structure makes it resistant to such unfolding and degradation, the researchers suggest.
This may not be an advantage for the organism as whole, however. “If knotted proteins are in fact more difficult to degrade, it might also be disadvantageous for most proteins to be knotted in the first place,” Virnau, Mirny, and Kardar say.
There’s much to be learned yet about the special role of knots in proteins. In this case, it’s not just a matter of spontaneous entanglement.