DARK MATTER Most microbes have stayed hidden from scientists, but new technologies are revealing unknown species of bacteria, some of which may hold medicinal promise.
Ashley Mackenzie
Few people today could recite the scientific accomplishments of 19th century physician Julius Petri. But almost everybody has heard of his dish.
For more than a century, microbiologists have studied bacteria by isolating, growing and observing them in a petri dish. That palm-sized plate has revealed the microbial universe — but only a fraction, the easy stuff, the scientific equivalent of looking for keys under the lamppost.
But in the light — that is, the greenhouse-like conditions of a laboratory — most bacteria won’t grow. By one estimate, a staggering 99 percent of all microbial species on Earth have yet to be discovered, remaining in the shadows. They’re known as “microbial dark matter,” a reference to astronomers’ description of the vast invisible matter in space that makes up most of the mass in the cosmos.
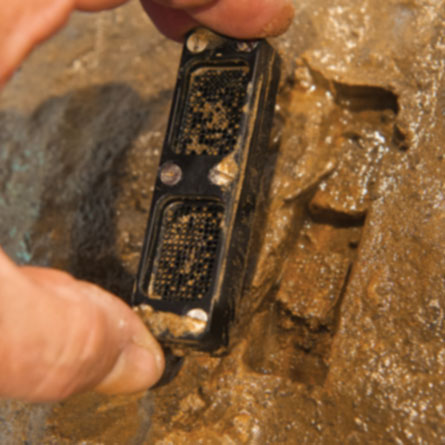

In the last decade or so, though, scientists have developed new tools for growing bacteria and collecting genetic data, allowing faster and better identifications of microbes without ever removing them from natural conditions. A device called the iChip, for instance, encourages bacteria to grow in their home turf. (That device led to the discovery of a potential new antibiotic, in a time when infections are fast outwitting all the old drugs.) Recent genetic explorations of land, water and the human body have raised the prospect of finding hundreds of thousands of new bacterial species.
Already, the detection of these newfound organisms is challenging what scientists thought they knew about the chemical processes of biology, the tree of life and the manner in which microbes live and grow. The secrets of microbial dark matter may redefine how life evolved and exists, and even improve the understanding of, and treatments, for many diseases.
“Everything is changing,” says Kelly Wrighton, a microbiologist at Ohio State University in Columbus. “The whole field is full of enthusiasm and discovery.”
Counter culture
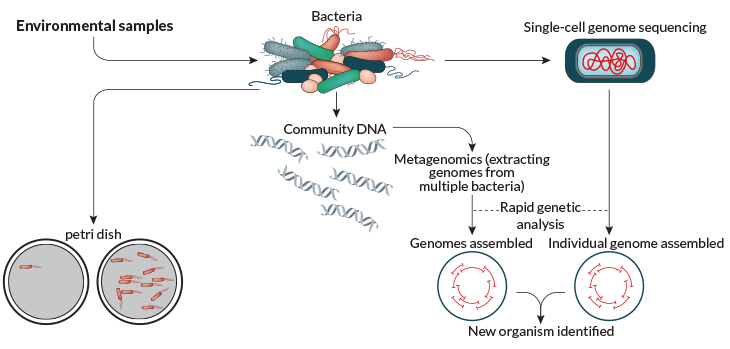
Microbiologists have in the past discovered new organisms without petri dishes, but those experiments were slow going. In one of her first projects, Tanja Woyke analyzed the bacterial community huddled inside a worm that lives in the Mediterranean Sea. Woyke, a microbiologist at the U.S. Department of Energy’s Joint Genome Institute in Walnut Creek, Calif., and colleagues published the report in Nature in 2006. It was two years in the making.
They relied on metagenomics, which involves gathering a sample of DNA from the environment — in soil, water or, in this case, worm insides. After extracting the genetic material of every microbe the worm contained, Woyke and colleagues determined the order, or sequences, of all the DNA units, or bases. Analyzing that sequence data allowed the researchers to infer the existence of four previously unknown microbes. It was a bit like obtaining boxes of jigsaw puzzle pieces that need assembly without knowing what the pictures look like or how many different puzzles they belong to, she says. The project involved 300 million bases and cost more than $100,000, using the time-consuming methods available at the time.
Just as Woyke was wrapping up the worm endeavor, new technology came online that gave genetic analysis a turbo boost. Sequencing a genome — the entirety of an organism’s DNA — became faster and cheaper than most scientists ever predicted. With next-generation sequencing, Woyke can analyze more than 100 billion bases in the time it takes to turn around an Amazon order, she says, and for just a few thousand dollars. By scooping up random environmental samples and searching for DNA with next-generation sequencing, scientists have turned up entirely new phyla of bacteria in practically every place they look. In 2013 in Nature, Woyke and her colleagues described more than 200 members of almost 30 previously unknown phyla. Finding so many phyla, the first big groupings within a kingdom, tells biologists that there’s a mind-boggling amount of uncharted diversity.
Woyke has shifted from these broad genetic fishing expeditions to working on individual bacterial cells. Gently breaking them open, she catalogs the DNA inside. Many of the organisms she has found defy previous rules of biological chemistry. Two genomes taken from a hydrothermal vent in the Pacific Ocean, for example, contained the code UGA, which stands for the bases uracil, guanine and adenine in a strand of RNA. UGA normally separates the genes that code for different proteins, acting like a period at the end of a sentence. In most other known species of animal or microbe, UGA means “stop.” But in these organisms, and one found about the same time in a human mouth, instead of “stop,” the sequence codes for the amino acid glycine. “That was something we had never seen before,” Woyke says. “The genetic code is not as rigid as we thought.”
Other recent finds also defy long-held notions of how life works. This year in the ISME Journal, Ohio State’s Wrighton reported a study of the enzyme RubisCO taken from a new microbial species that had never been grown in a laboratory. RubisCO, considered the most abundant protein on Earth, is key to photosynthesis; it helps convert carbon from the atmosphere into a form useful to living things. Because the majority of life on the planet would not exist without it, RubisCO is a familiar molecule — so familiar that most scientists thought they had found all the forms it could take. Yet, Wrighton says, “we found so many new versions of this protein that were entirely different from anything we had seen before.”
The list of oddities goes on. Some newly discovered organisms are so small that they barely qualify as bacteria at all. Jillian Banfield, a microbiologist at the University of California, Berkeley, has long studied the microorganisms in the groundwater pumped out of an aquifer in Rifle, Colo. To filter this water, she and her colleagues used a mesh with openings 0.2 micrometers wide — tiny enough that the water coming out the other side is considered bacteria-free. Out of curiosity, Banfield’s team decided to use next-generation sequencing to identify cells that might have slipped through. Sure enough, the water contained extremely minuscule sets of genes.
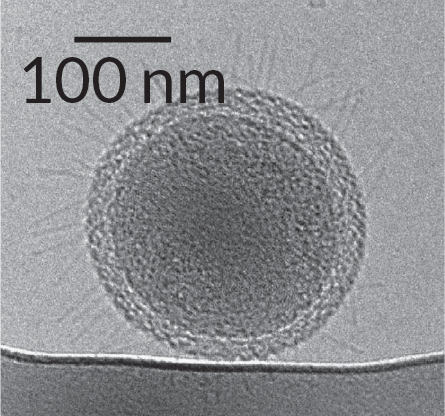
“We realized these genomes were really, really tiny,” Banfield says. “So we speculated if something has a tiny genome, the cells are probably pretty tiny, too.” And she has pictures to prove it. Last year in Nature Communications, she and her team published the first images (taken with an electron microscope) and detailed description of these ultrasmall microbes (see, right). They are probably difficult to isolate in a petri dish, Banfield says, because they are slow-growing and must scavenge many of the essential nutrients they need from the environment around them. Part of the price of a minigenome is that you don’t have room for the DNA to make everything you need to live.
Relationship status: It’s complicated
Banfield predicts that an “unimaginably large number” of species await in every cranny of the globe — soil, rocks, air, water, plants and animals. The human microbiome alone is probably teeming with unfamiliar microbial swarms. As a collection of organisms that live on and in the body, the human microbiome affects health in ways that science is just beginning to comprehend (SN: 2/6/16, p. 6).
Scientists from UCLA, the University of Washington in Seattle and colleagues recently offered the most detailed descriptions yet of a human mouth bacterium belonging to a new phylum: TM7. (TM stands for “Torf, mittlere Schicht,” German for a middle layer of peat; organisms in this phylum were first detected in the mid-1990s in a bog in northern Germany.) German scientists found TM7 by sifting through soil samples, using a test that’s specific for the genetic information in bacteria. In the last decade, TM7 species have been found throughout the human body. An overabundance of TM7 appears to be correlated with inflammatory bowel disease and gum disease, plus other conditions.
Until recently, members of TM7 have stubbornly resisted scientists’ efforts to study them. In 2015, Jeff McLean, a microbiologist at the University of Washington, and his collaborators finally isolated a TM7 species in a lab and deciphered its full genome. To do so, the team combined the best of old and new technology: First the researchers figured out how to grow most known oral bacteria together, and then they gradually thinned down the population until only two species remained: TM7 and a larger organism.
“The really remarkable thing is we finally found out how it lives,” McLean says, and why it wouldn’t grow in the lab. They discovered that this species of TM7, like the miniature bacteria in Colorado groundwater, doesn’t have the cellular machinery to get by on its own. Even more unusual, these bacteria pilfer missing amino acids and whatever else they need by latching on, like parasites, to a larger bacterium. Eventually they can kill their host. “We think this is the first example of a bacterium that lives in this manner,” McLean says.
He expects to see more unusual relationships among microbes as the dark matter comes to light. Many have evaded detection, he suspects, because of their small size (sometimes perhaps mistaken for bacterial debris) and dependence on other organisms for survival. In 2013 in the Proceedings of the National Academy of Sciences, McLean and colleagues were the first to describe a member of another uncultivated phylum, TM6. They found this group growing in the slime in a hospital sink drain. Later studies determined that the organism lives by tucking itself inside an amoeba.
One of the greatest hopes for microbial dark matter exploration is that newly found microbes might provide desperately needed antibiotics. From the 1940s to the 1960s, scientists discovered 10 new classes of drugs by testing chemicals found in soil and elsewhere for action against common infections. But only two classes of medically important antibiotics have been discovered in the last 30 years, and none since 1997. Some major infections are at the brink of being unstoppable because they’ve become resistant to most existing drugs (SN Online: 5/27/16). Many experts think that natural sources of antibiotics have been exhausted.
Maybe not. In 2015, a research team led by scientists from Northeastern University in Boston captured headlines after describing in Nature a new chemical extracted from a ground-dwelling bacterium in Maine. The scientists isolated the organism using the iChip, a thumb-sized tool that contains almost 400 separate wells, each large enough to hold only an individual bacterial cell plus a smidgen of its home dirt. The bacteria grow on this scaffold in part because they never leave their natural surroundings. In lauding the discovery, Francis Collins, director of the National Institutes of Health, called the iChip “an ingenious approach that enhances our ability to search one of nature’s richest sources of potential antibiotics: soil.” So far, the research team has discovered about 50,000 new strains of bacteria.
One strain held an antibiotic, named teixobactin (SN: 2/7/15, p. 10). In laboratory experiments, it killed two major pathogens in a way that did not appear easily vulnerable to the development of resistance. Most antibiotics work by disrupting a microbe’s survival mechanism. Over time, the bacteria genetically adapt, find a work-around and overcome the threat. This new antibiotic, however, prevents a microbe from assembling the molecules it needs to form an outer wall. Since the antibiotic interrupts a mechanical process and not just a specific chemical reaction, “there’s no obvious molecular target” for resistance, says Kim Lewis, a microbiologist at Northeastern.
Everything is illuminated
Some microbiologists feel like astronomers who, after years of staring up into the dark, were just handed the Hubble Space Telescope. Billions of galaxies are coming into view. Banfield expects this new microbial universe to be mapped over the next few years. Then, she says, an even more exciting era begins, as science explores how these dark matter bacteria make a living. “They are doing a lot of things, and we have no idea what,” she says.
Part of the excitement comes from knowing that microbes have a history of granting unexpected solutions to problems that scientists never expected to solve. Consider that the enzyme that makes the laboratory technique PCR possible came from organisms that live inside the thermal vents at Yellowstone National Park. PCR, which works like a photocopier to make multiple copies of DNA segments, is now used across a range of situations, from diagnosing cancer to paternity testing. CRISPR, a powerful gene-editing technology, relies on “molecular scissors” that were found in bacteria (SN: 9/3/16, p. 22).
Banfield estimates that 30 to 50 percent of newly discovered organisms contain proteins that never met a petri dish. Their function in the chemistry of life is an obscure mystery. Since microbes are the world’s most abundant organism, Banfield says, “the vast majority of life consists of biochemistry we don’t understand.” But once we do, the future could be very bright.
This article appears in the September 17, 2016, issue of Science News with the headline, “Out of the dark.”