DIVERSE DNA A new database containing information about DNA variants in protein-coding genes of more than 60,000 people may be a valuable resource for finding the causes of genetic diseases.
gecko753/istockphoto
A new catalog of human genes reveals that people have many different ways to build proteins. This listing of options can help doctors sort through mutations to learn which ones cause genetic diseases — and which ones don’t.
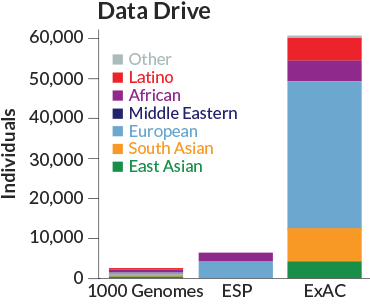
An international group of researchers banded together to compile the catalog, an inventory of the exome — the small portion of the human genome that produces proteins — of 60,706 adults from different populations around the world. Researchers in the Exome Aggregation Consortium, known as ExAC, report the findings online October 30 at bioRxiv.org.
“This is one of the most useful resources ever created for medical testing for genetic disorders,” says Heidi Rehm, a clinical lab director at Harvard Medical School who is not a member of the consortium.
A journal reviewing the work for publication prohibits the ExAC researchers from speaking with journalists about the manuscript posted on bioRxiv.org, one researcher involved in the project told Science News. The work has yet to be peer-reviewed and researchers are not allowed to “publicize” their findings before they have been vetted by their peers. Other researchers have already viewed the manuscript on bioRxiv.org and pointed out a few minor flaws, including broken links and formatting errors. No one has yet criticized the data or analysis.
“This work is both technically very impressive … and will be a fantastic mine of information to explore over the next years, and also hugely useful in clinical genetics settings,” says Gilean McVean, a statistical geneticist at the University of Oxford. Looking at just the protein-coding parts of the genome is a good start, he adds, “but we will need the full spectrum of the whole genome to ultimately make sense of what causes disease.”
Among the people who donated DNA to the project, the ExAC researchers found more than 7.4 million genetic variants, letters in the DNA instructions for building proteins that differ from one person to another. On average, people had one genetic variant for every eight base pairs, the information-carrying chemicals that make up DNA.
Those variants aren’t spread evenly among genes, though. The researchers found that 3,230 genes are almost devoid of any harmful variants. That finding provides “an exquisitely detailed view into what genetic perturbations are ‘biologically permissible,’” McVean says.
Genes that don’t have mutations are likely to be ones important for human development and survival, says Rehm. Such genes, when mutated, may cause severe genetic disease or stop an embryo from developing so no living person would carry mutations in those genes.
For other genes, “lightning does strike several times in the same spot,” says Tuuli Lappalainen, a geneticist at the New York Genome Center and Columbia University. About 43 percent of new mutations in a child that are not also present in the parents turned out to be copycats of variants carried by other people in the ExAC database. That means that doctors who just look for new mutations to explain a child’s genetic disease could mistake these types of mutations for disease-causing ones even though they are harmless.
The exome project may help medical geneticists avoid making similar mistakes due to not knowing how rare variants are. The data have revealed that some variants are not as uncommon as researchers previously believed. For instance, some variants rarely show up in some populations, but are relatively common in people from another part of the world. Finns are a good example: Finland had a small founding population so some mutations are found in Finns more frequently than in other Europeans.
In addition, an average participant in the exome project harbors about 53 variants that have previously been classified as disease-causing. But, on average, 41 of those mutations are found relatively frequently in at least one population, where they do not cause disease, the data show.
The ExAC team discovered an example of such false accusation when it investigated 192 variants that had previously been implicated in disease. These variants were rarely found in people in the limited datasets researchers could access before. But the ExAC data show that many of those variants are found in more than 1 percent of healthy South Asian or Latino people, indicating that they are probably not the culprits. The misleading mutations include a variant thought to cause a liver disease known as North American Indian Childhood Cirrhosis when children inherit two copies of the variant. That variant was found in 226 Latin Americans, including four people who had two copies of the gene but didn’t have the liver disease. That result suggests that the variant isn’t the source of the liver disease.
Researchers outside the ExAC team have had access to the data for more than a year, but those scientists have agreed not to publish large-scale findings until after the ExAC team reports their methods and analysis in a scientific journal. Lappalainen says she expects ExAC’s official debut to be accompanied by multiple companion papers, followed by researchers using the data in other types of studies. Those studies may guide doctors toward better diagnosis of genetic diseases and suggest treatments.
Already, thousands of patients may need to have their cases reevaluated in light of the new data, Rehm says.
Editor’s Note: This story was updated November 20, 2015, to correct the description of participants in the study. People with diseases were included, so not all were healthy.