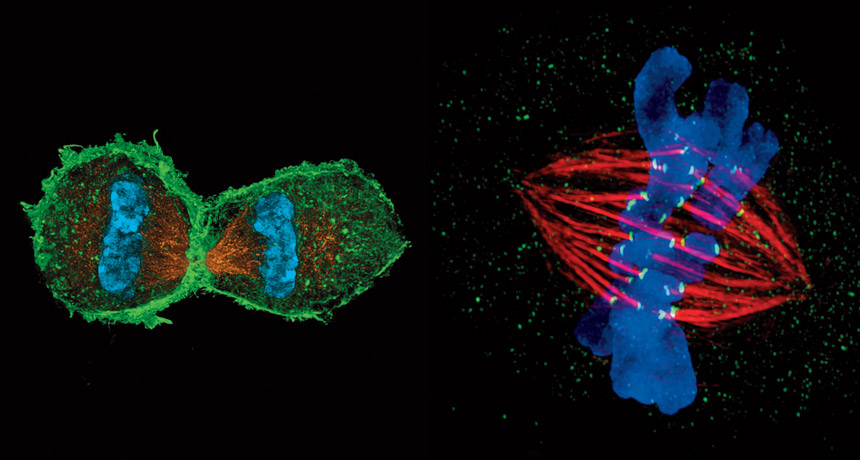
Left: L. Schermelleh/Univ. of Oxford; Right: J. Stout/Indiana Univ.
Imagine if your best knowledge of human anatomy came from viewing the body through binoculars from a mile away. You might make out the shape of a hand, but knuckles and fingernails would elude you. Experiments could tell you there’s a pumping heart inside, but to see that heart with any clarity you would have to fix it in formaldehyde or liquid nitrogen, blast it with electrons and add dyes to impart contrast.