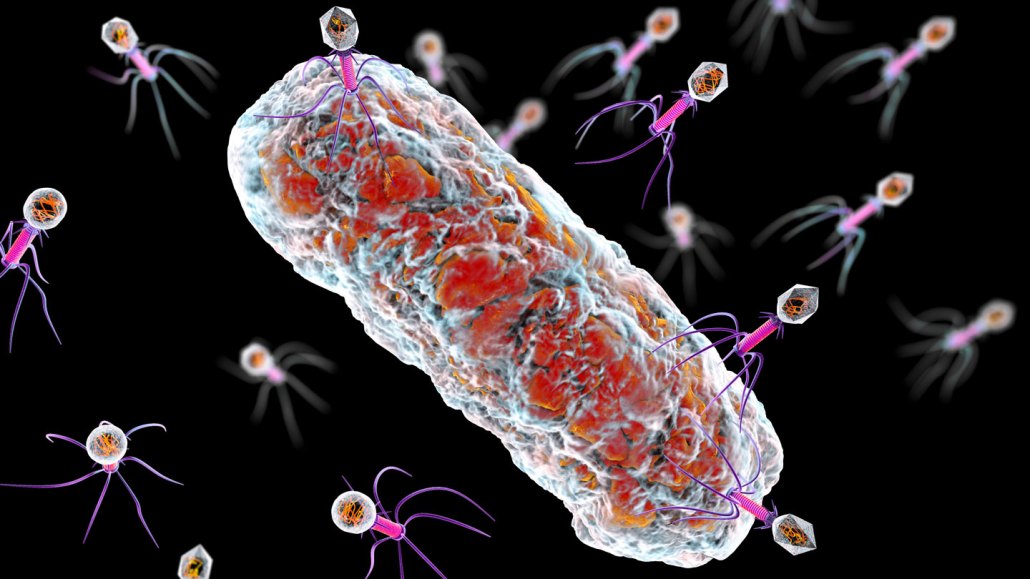
When bacteria evolve to become resistant to bacteria-killing viruses known as bacteriophages (illustrated attacking a bacterium), it doesn’t always end well for the bacteria.
Kateryna Kon/Science Photo Library/Getty Images Plus

When bacteria evolve to become resistant to bacteria-killing viruses known as bacteriophages (illustrated attacking a bacterium), it doesn’t always end well for the bacteria.
Kateryna Kon/Science Photo Library/Getty Images Plus