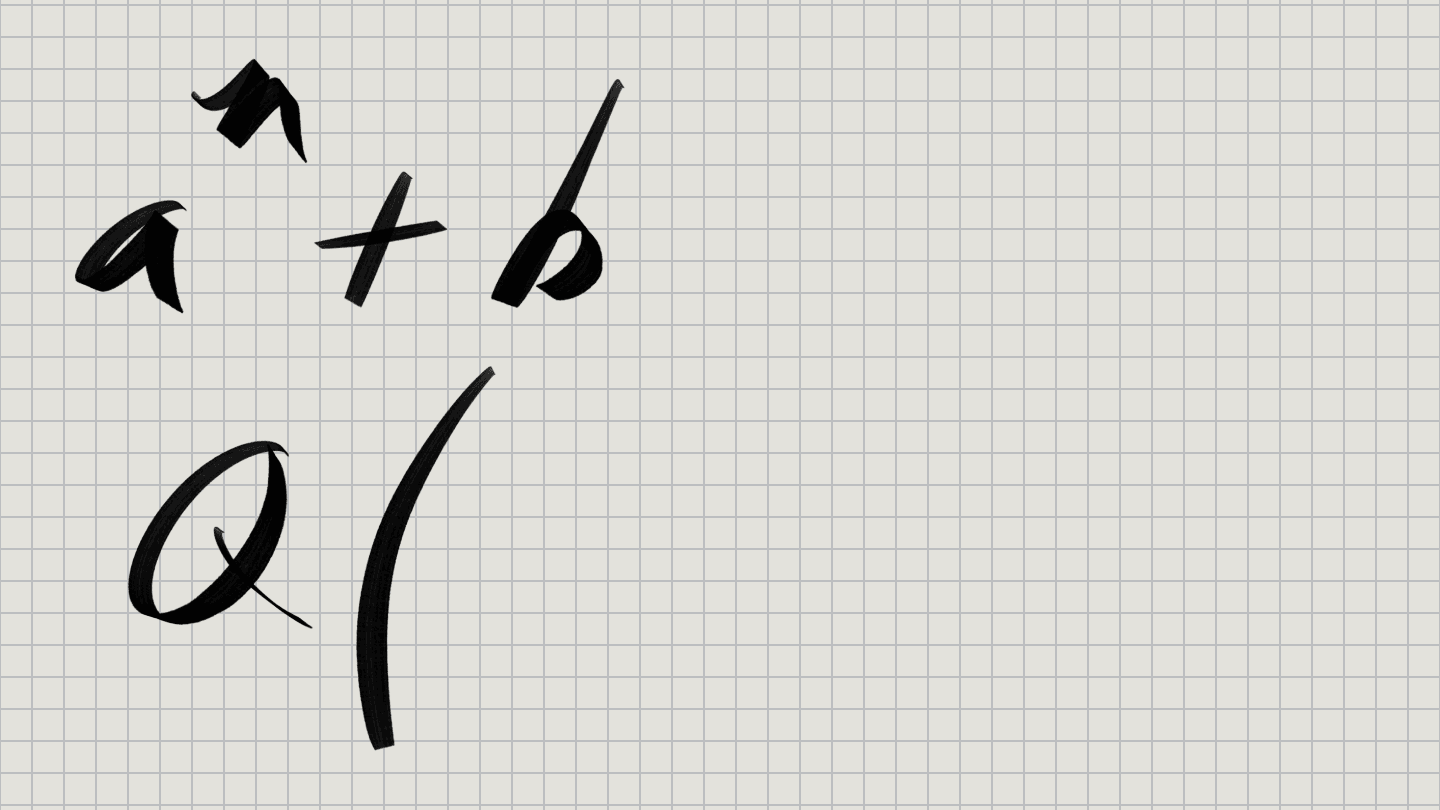Detangling DNA
DNA can form some very nasty knots — but not just any knots
- More than 2 years ago
Deep inside our cells, the DNA that encodes the mysteries of our individuality twines into tidy little spiral staircases neatly side by side — or so we might imagine.
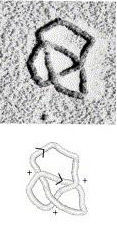
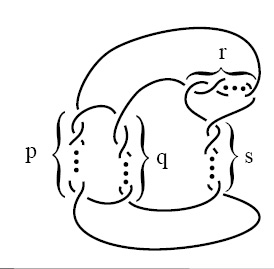
Consider, though, that if you scale up the nucleus of a cell to the size of a basketball, each molecule of DNA inside it would resemble fishing line more than four miles long. And now consider what happens to your iPod headphones when you cram them into a pocket: Invariably, it seems, they tangle. And they’re only a foot long!
Now you have a picture of the gargantuan task your cells face in managing the snarls that form in DNA. Storing the DNA isn’t a problem because the cell can pack each strand systematically into a tidy, tight ball. And for some tasks, the cell can just unwind the ball a bit, keeping the unruly strands in check. But when the cell needs to snip the DNA and rearrange its genetic sequence, the strands almost unavoidably kink into a tangled mess.
Researchers have found that DNA can form incredibly complex knots, sometimes with dozens of crossings. But now a pair of mathematicians has shown that DNA can only form certain kinds of knots, not any knot at all. The discovery may help biologists understand site-specific recombination, the way that cells perform surgery on their DNA.
Although we tend to think of our genetic sequence as being fixed at conception, cells occasionally need to shuffle specific bits of their DNA around. Cells might reverse a small stretch of a sequence or move a section from one strand to another. Brewer’s yeast, for example, uses recombination just before cell division to prepare its DNA to divide rapidly. Viruses use recombination to insert their own DNA into the host cell, tricking the cell into producing thousands of copies of the virus. And recombination is our tool when we create genetically engineered cells.
But the process almost always causes some nasty knots. To alter the genetic sequence, specialized enzymes grab two pieces of DNA, snip them apart, bring the ends together, reshuffle the genetic sequence between them and rejoin the ends. Because the DNA is so tightly packed together, this process often ends with strands wound around themselves or one another, forming a knot or link. Another type of enzyme cleans up the mess afterward, snipping strands, passing them through others and reattaching them until the knot is unwound.
Biologists still don’t understand very well how recombination works. “What you really want is to see an enzyme attaching to the DNA and watch it dragging it around,” says Dorothy Buck, a mathematical biologist at Imperial College London. “You want a YouTube video of the whole process, but you can’t get it.” Current technology in microbiology only allows for still images. And even getting still images of knotted DNA is very tricky.
Biologists have managed to learn some things about how recombination works, though. Buck and her collaborator Erica Flapan of PomonaCollege in Claremont, Calif., found three rules governing the behavior of the enzymes that perform the recombination. The precise path the enzymes take and the way they perform their surgery determines the knot that is formed. Buck and Flapan realized that their rules meant that site-specific recombination could twist the DNA only into particular knots and links, and they applied knot theory to figure out which ones they were. Only a small proportion of the very complicated knots could occur in DNA, they found.
Narrowing down the possible knots could in turn shed light back on the activities of the enzymes. Ultimately, this could help researchers use site-specific recombination to repair the faulty DNA that causes genetic diseases.