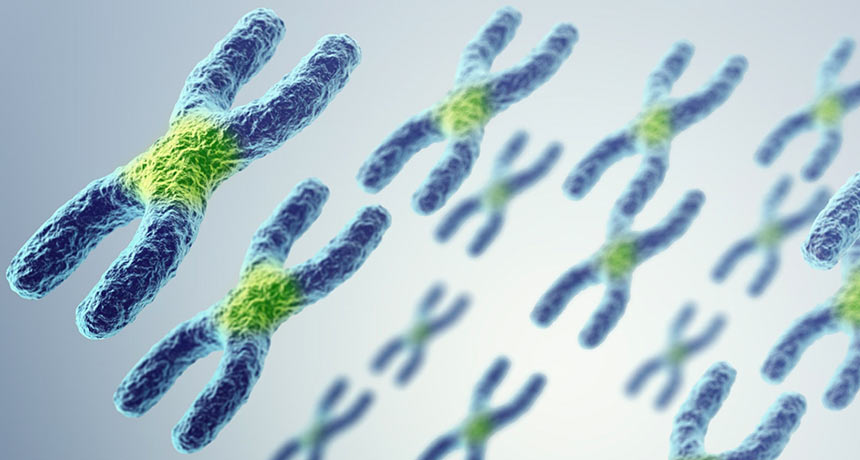
CORE STRENGTH Researchers have decoded the genetic composition of important structures called centromeres (colored green in this illustration) in fruit flies’ chromosomes. Centromeres ensure that chromosomes are properly distributed to dividing cells.
Rost-9D/iStock / Getty Images Plus
Some of the most important chapters in fruit flies’ genetic instruction book have finally been decoded.
For the first time, researchers have deciphered, or sequenced, the genetic makeup of all of a multicellular organism’s centromeres — and discovered stretches of DNA that may be key in divvying up chromosomes. Errors in doing that job can lead to cancer, birth defects or death. The team reported the achievement May 14 in PLOS Biology.
Centromeres, which give most chromosomes their characteristic X shape, help move chromosomes in dividing cells. “The chromosome is a bus, and our DNA and genes are the passengers. The centromere is the bus driver,” says Beth Sullivan, a geneticist and centromere biologist at Duke University School of Medicine not involved in the study. “It’s what moves the chromosome, after the DNA has been copied, into new daughter cells.”
Until now, scientists have known very little about these genetic bus drivers. Some centromeres from corn, horses, yeast and other fungi — and one human one that drives the Y chromosome — have been characterized. But mostly what scientists knew about centromeres is that they are incredibly long stretches of repetitive DNA.

The reason for the oversight was technical: To sequence the genome — meaning determine the order of DNA’s chemical letters, known as bases — scientists first had to chop the DNA into tiny pieces about 150 bases long. Computers can put the genome together by finding where two pieces match up. Centromeres tend to repeat the same letters over and over again and can stretch for millions of bases, so many of the small bits look alike. “It’s like trying to put together a jigsaw puzzle of a blue sky when all the puzzle pieces look the same,” says Amanda Larracuente, an evolutionary geneticist at the University of Rochester in New York.
Larracuente and colleagues used new technology to sequence stretches of DNA 10,000 to 100,000 bases long. “It’s like having much bigger puzzle pieces,” Larracuente says, “So now we might be able to catch a wisp of a cloud in our blue-sky puzzle pieces, and that makes it a lot easier to figure out how these pieces of DNA fit together.” The team also used several other techniques to fit their centromere puzzles together.
Those techniques should make it possible for other scientists to fill in gaps in other organisms’ genomes, says Gernot Presting, a centromere biologist at the University of Hawaii at Manoa not involved in the study.
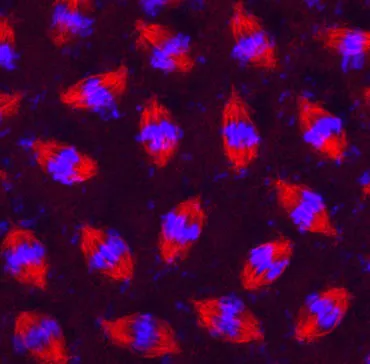
Each fruit fly chromosome’s centromere is unique, although all have the same basic structure of retrotransposon islands — particularly the retrotransposon G2/Jockey-3 — surrounded by seas of repetitive DNA. Copies of G2/Jockey-3 are scattered in other places in the fruit fly genome, but about 63 percent of its copies are in centromeres, the researchers found. Those results might mean that the transposon is a seed around which new centromeres can form, or the transposon may just hop into centromeres more often than into other parts of the genome.
The researchers uncovered one clue hinting that G2/Jockey-3 might seed centromeres: A key centromere protein called CENP-A clings to those transposon islands. That protein helps secure the centromere to the cellular machinery the separates chromosomes into new daughter cells. Finding that CENP-A sticks to the transposon islands suggests the jumping genes might be in the centromere’s driver’s seat.
Centromeres in plants, fungi, bats, gibbons and humans also contain retrotransposons. But no one really knows whether those transposons are driving the bus or are just along for the ride, says coauthor Barbara Mellone, a geneticist at the University of Connecticut in Storrs. Presting, for example, argues that there’s evidence that transposons are passengers: In corn, centromeres move first and retrotransposons hop aboard later. Armed with centromeres’ structure, Mellone and colleagues hope to find out exactly how important the jumping genes are to keeping fruit fly centromeres and their chromosomes on the right track.
Editor’s note: This story was updated May 17, 2019, to clarify the headline. While centromeres are some of the final parts of the fruit fly’s genetic makeup to be decoded, they aren’t the final bit.