“RESISTANT” shouts the title of Lindsay Richman’s post. Apparently, she was elated to learn that her DNA reduces her susceptibility to norovirus infections, the principal cause of the common stomach flu.
So she posted a comment on a discussion board on the website for 23andMe, a company based in Mountain View, Calif., that specializes in the fledgling industry of personal genomics. To get a glimpse of her own DNA, Richman had sent the company $400 and a vial of her spit. From her point of view, what happened next was a mystery — a black box. But a few weeks later, out popped her results on a password-protected website, complete with social networking tools for sharing and discussing her genetic inheritance with other customers.
In the string of online responses to Richman’s post, others who share her genetic good fortune compared notes on the last time they’d had any symptoms of stomach flu. Richman, a 26-year-old real estate agent in New York City, hasn’t had stomach flu since she was 8, she wrote.
In other discussions, people compared their genetic profiles and brainstormed on how lifestyle choices and environmental exposures might influence their risks for various conditions, such as Parkinson’s disease and prostate cancer.
Now that prices charged by personal genomics companies such as 23andMe, Navigenics, deCODE genetics and DNA Direct have dropped, ranging from a few hundred to a few thousand dollars, many people curious about their genetic inheritance, and how it relates to their health, have easier access to DNA testing.
Serving to “crowdsource” the search for new links among genes, behavior and disease, these companies’ customers represent a small army of amateur genome sleuths who could prove to be a new force in pushing genomic research forward. But the genetic report cards these amateurs are reading may not be as definitive as they assume. Despite progress in linking genetic differences with disease risk and other traits, the predictive power of these links has fallen short of expectations.
In April, the New England Journal of Medicine published a review and a set of essays grappling with this shortfall in DNA’s predictive power and searching for the best way to take research forward. An essay by Peter Kraft and David Hunter, epidemiologists at the Harvard School of Public Health in Boston, was revealingly titled, “Genetic risk prediction — are we there yet?”
Their answer, in a nutshell: No. Which leads to the crucial question of what, exactly, customers of personal genomics companies are looking at when logging on to the digital oracle to peek at their genetic fates.
The leap from a tube of saliva to a ledger of traits, health risks and ancestral history involves a lot of science — some credible, some flimsy. As more and more people become consumers of genetic information services, these people may want to first open that black box and take a good look inside.

The black box
It’s certainly not the most dignified way to join the genomics era.
Inside the small, brightly colored DNA-sample kit that arrives in the mail lies a clear plastic tube capped by a blue plastic funnel — the easier to spit into. Users are instructed to fill the tube to a little line, which the directions say can take five minutes or more. Five minutes of repetitive spitting.
In that saliva float cheek cells that have sloughed off from the soft tissue lining the mouth. Snapping the tube’s lid shut releases a preservative that keeps the cells intact during their voyage to the lab.
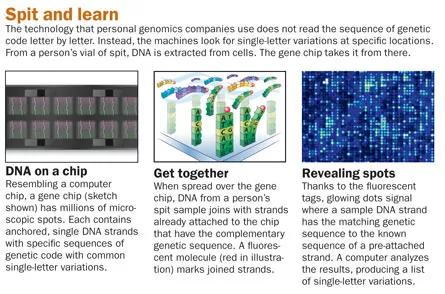
“The reason we collect so much spit is to get a lot of your DNA,” explains Brian Naughton, a founding scientist at 23andMe. The machines that read the sequence of DNA chemical “letters” of the genetic code are quite accurate and robust against noise. Repeat the scan with DNA from the same person, and the two results will be more than 99.9 percent identical, according to the company.
“It’s clearly reproducible,” says George M. Church, a geneticist at Harvard Medical School in Boston. Testing a person again with another company’s service using a different model of DNA reading machine will also produce nearly identical results, Church says.
These machines use gene chips to read up to a million letters of genetic code at once. Since the first gene chips that could skim the entire genome debuted in 2005, the cost to scan a person’s DNA has dropped dramatically.
Despite what some customers might assume, most personal genomics companies do not produce a complete sequence of a customer’s genome. An entire human genome contains about 6 billion letters of genetic code distributed among a person’s 23 pairs of chromosomes (the inspiration for the name 23andMe). These machines don’t read every DNA letter. Instead, they read individual letters at 500,000 to a million different spots in the genome, capturing just 8 to 16 thousandths of one percent of the full genome.
But that genome sliver is carefully selected to represent much of the genetic variation among people. “It’s really coverage of that genetic variation that you’re going for,” Naughton explains.
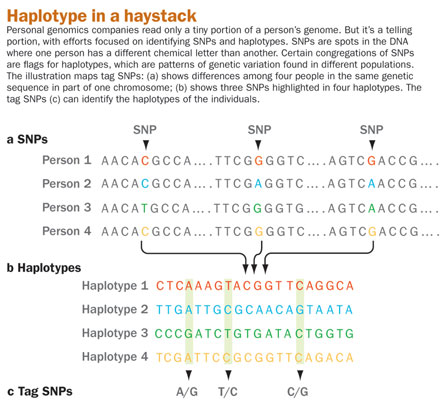
Nearly all of a person’s genetic code is identical to that of every other person — roughly 99.5 percent of it matches up letter for letter. At certain spots along the length of a person’s chromosomes, though, the genetic code can differ from other people’s by a single letter. In the sequence of the familiar A’s, T’s, C’s and G’s, the four information-carrying nucleo- tides in DNA, some people might have a T at a certain location while others have a C. These small variations are called single nucleotide polymorphisms, or SNPs (pronounced “snips”).
The human genome contains about 10 million known SNPs. That estimate is the latest from the International HapMap Project, an ongoing scientific collaboration that’s mapping these genetic variations. But SNPs near each other on a chromosome tend to get inherited together, making it possible to group neighboring SNPs into units of inheritance called haplotypes. A few tag SNPs are enough to identify each group, so it takes only about 500,000 tag SNPs to reveal a person’s haplotypes. That’s why most gene chips test for at least 500,000 SNPs, and why checking this tiny sliver of the genome — these tag SNPs — can reveal much of the genetic variation that makes a person unique.
Tag SNPs usually aren’t part of any known gene. But the surrounding DNA inherited along with a SNP can contain one or more genes important for various diseases. The SNP is often a proxy.
This much of the black box is fairly reliable: the raw data on at least hundreds of thousands of a person’s SNPs. Some personal genomics companies make this mountain of raw data directly available to their customers.
But that’s the easy part. The other, more problematic half of the black box is interpretation. What exactly does having one SNP variant or another mean for a person’s risk for heart disease, diabetes, colon cancer?
In short, there’s no single answer for how reliable these interpretations are. Some SNPs have clear, strong and well-understood links with specific traits or diseases. Many others have only small effects, and the biological mechanisms for the links are often unknown.
Reviewing the available scientific evidence for each disease or trait is the biggest challenge for these companies, says Michele Cargill, director of human genetics for Navigenics. “You really have to read each [study] very, very carefully. It takes a lot of time and it’s all done manually.”
For example, the resistance to norovirus infection that Richman enjoys is linked to a SNP called rs601338, which is located on each of the two chromosome 19s she inherited from her parents. At this location, a person can have either an A or a G for this SNP. Chromosomes with an A lack a functioning copy of a gene called FUT2. This gene produces a certain molecule on the outer surfaces of the cells that line the intestines. As it turns out, noroviruses must bind to this molecule in order to enter and infect the cells. People like Richman, who inherited the A SNP from both of her parents, do not have a working copy of the FUT2 gene, so their intestinal cells lack the molecule that noroviruses need to cause an infection.
In this case, the link is strong. A single SNP can indicate whether a person has a working copy of a certain gene, and the biological mechanisms tying this gene to the virus’s ability to enter the person’s cells are well understood. In nearly all studies, people who lacked a functional copy of the FUT2 gene didn’t get sick, even when deliberately exposed to this type of virus. It’s as close as one ever gets in biology to a slam dunk.
About 1,300 genes have known, strong links to medical conditions, Church notes. Some disorders, such as sickle-cell anemia, are truly genetic diseases that a person has from birth. Others such as Parkinson’s disease, macular degeneration, Alzheimer’s disease and breast cancer arise later in life and can be influenced by environment and lifestyle, even though some genes are known to significantly change the odds that a person will get the disease.
But the relatively clear-cut cases are the exception, not the rule. For many traits and diseases, finding reasonably strong and reliable links with SNPs has proven more difficult than many scientists had expected.
Beware weak links
To search for these links, researchers use gene chips similar to those used by personal genomics companies. With these gene chips, scientists scan the DNA of two groups of people: a few hundred or thousand people with the disease in question and a few hundred or thousand without it. If the two groups are well matched in terms of other important traits such as age, ethnicity, smoking habits and so on, a SNP that appears more frequently among people with the disease than among the control group may be associated with the disease.
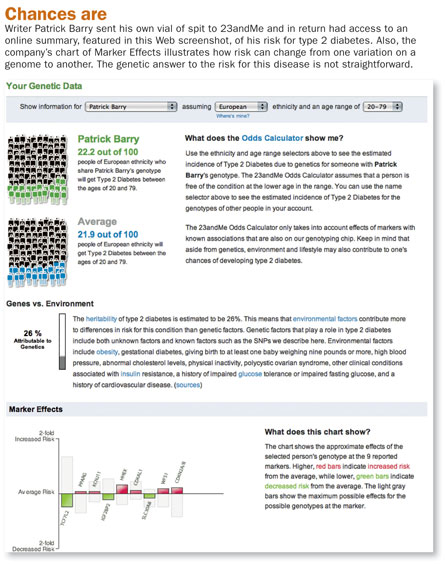
At first, these studies tended to “discover” a lot of illusory links — false connections that crop up by pure chance in the mountains of data produced by these studies (SN: 6/21/08, p. 20). In the last few years, scientists have learned to correct for these statistical sins, but even with most false positives weeded out, these studies often point to large numbers of potentially suspect SNPs. For example, dozens of well-established SNPs contribute to the risk for type 2 diabetes, the form of the disease that emerges in late adulthood and is hastened by a diet high in sugar.
Unfortunately, most of these SNPs alter risk by only a tiny amount — a 2 percent change in risk for this one, a 5 percent change for that one, a 10 percent change for a third. Even a 10 percent change would mean that the odds of developing diabetes sometime in that person’s lifetime would increase from the typical 22 percent to 24 percent. Not exactly a crystal ball–caliber revelation.
“People will tell you that if they know that they have a slightly higher risk for diabetes, then they could change their diet,” Kraft says. “But you don’t really need an expensive genome test to tell you that.”
Even more problematic, though, is the fact that new links continue to trickle in. When the risks from individual SNPs are added, a person’s overall risk of the disease could be slightly positive. But a newly discovered SNP could tilt the balance in the other direction.
“Our best guess about your risk today might turn out to be very different five years from now,” Kraft says. “Your risk as a function of time is a random walk. It bounces up and down as we learn more.”
These problems also plague other common illnesses such as heart disease, as well as complex traits such as height and intelligence. In each of these cases, large numbers of SNPs are involved because the underlying biology is complicated. Heart disease, for example, is an umbrella term for various cardiovascular problems that could depend on genes for heart muscle proteins, blood vessel strength and elasticity, cholesterol metabolism, blood clotting and others. And each of these functions often arises from webs of interactions among dozens or hundreds of genes. These interactions produce the feedback loops that fine-tune a cell’s behavior and make it robust, and these interactions add to genetic complexity (SN: 12/6/08, p. 22).
“It’s very early days in terms of what we’ve learned about common, complex diseases,” Kraft says.
Some in the genome research community say that studies with ever larger numbers of subjects are needed to find SNPs that have even weaker links but that many people have. The hope is that the cumulative effects will point to significant predictions. Others suggest that studies have already found all the important, common SNPs that there are to find, and that new studies should instead search for SNPs that are held by a small minority of people but that exert a stronger influence on disease.
Selling, knowing risk
Until more progress is made, some scientists say, selling risk information about complex diseases is premature.
“The justification is just not there for claiming that these [SNPs] have any clinical utility at this stage” for complex diseases, says Allan Balmain, a cancer geneticist at the University of California, San Francisco. He says that, in his opinion, personal genomics companies are “just exploiting the naïveté in the general population.”
These companies plan to use their growing databases of DNA samples — and their legions of curious customers — for novel research. Other research scientists are cautiously optimistic. Customers who have sent their DNA samples to these companies are not a random sample of the population, a fact that could bias the results. And information about family history, health habits and environmental exposures are gathered through unmonitored online surveys, rather than by professional clinicians. With careful study design and quality control for the data, though, these obstacles could be surmountable. “Their data set is not in itself bad,” Balmain says. “Some of these things may turn out to be very useful.”
Eventually, though, these companies may need to move beyond SNPs, as many researchers have begun to do. Much of the genetic variation among people comes in forms other than changes to single letters of code. Some people have long chunks of DNA within their genomes that other people lack. Along with these insertions and deletions, the number of copies of some genes varies from person to person. While these kinds of structural differences are far less numerous than SNPs, each of them can involve hundreds or thousands of letters of genetic code, so together they account for about four times more genetic variation than SNPs do (SN: 4/25/09, p. 16).
Modified gene chips can detect some small insertions and deletions, but the best way to tally these changes is with the more thorough, and more expensive, approach of DNA sequencing.
Current state-of-the-art technologies can sequence an entire human genome for about $5,000, compared with millions of dollars just four years ago. “I think nobody anticipated just how fast the cost of sequencing would change,” Church says. “I think we’re already at the tipping point. It’s already getting feasible to sequence large portions of genomes, maybe all the coding regions, for studies and for individuals.”
Unfortunately, Naughton says, 23andMe doesn’t store frozen saliva samples for most customers, only for those participating in research projects. So if Richman someday wants to upgrade to a full genome sequence to learn even more about her genetic inheritance, she’ll have to go through all that spitting again.