In Silico Medicine
Computer simulations aid drug development and medical care
Like millions of people in the United States, Bill and Allen have asthma. They’re lucky enough to take the newest therapies, sometimes even before the drugs come to market. Yet neither Bill nor Allen has ever been to a doctor’s office or the hospital. After all, it’s okay if they get sick or even die–a simple click of a mouse can restore these patients to perfect health. Bill and Allen are two of the newest subjects in a growing research area called in silico biology. As basic researchers, drug developers, doctors, and health-care administrators struggle with the sheer volume and complexity of scientific information that they face every day, information gathered from computer-generated patients like Bill and Allen may lead to better decisions.

Computer simulations aren’t new in nonmedical research and development. Much of climatology and geology is modeled with computers, and engineers routinely design products–from airplanes to tire treads–using mathematical simulations rather than building and testing prototypes. But such computer applications have been slow to reach people studying the vagaries of the human body. Today, however, researchers are increasingly turning to computers to explore medical science.
“Computer simulation is the modus operandi for the future of biomedical research,” says James B. Bassingthwaighte of the University of Washington in Seattle. “The complexity of [biological] systems is such that intuition is of no help in figuring out where an intervention might have an effect or what is the most efficient means of diagnosis and treatment.”
Developing drugs
The company that Bill and Allen call home created its model of asthma patients by compiling information from more than 3,500 scientific studies on the disease. Next, scientists at the company–Entelos in Menlo Park, Calif.–wrote equations that account for more than 7,500 parameters of a person’s health that may be important in asthma. These include the thickness of the mucous lining in the lungs and the effects of inflammation in blocking the airways.
From one to the next, virtual patients may respond differently to drugs depending on their health parameters, just as real patients do, and their diseases may take different courses. But because the researchers know the exact differences between their virtual patients, they can more easily tease out the most important factors in complicated interactions.
Virtual patients may also respond differently when the modelers use different views of how a disease develops. Bill and Allen, for example, represent two hypotheses about how asthma attacks begin. Bill reflected the idea that the immune-signaling molecule interleukin-5 overstimulates immune cells called eosinophils. Those cells then trigger inflammation and asthma attacks. Allen, in contrast, had an asthma attack when immune cells called macrophages blocked his airways.
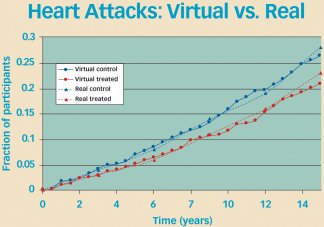
Entelos developed Bill and Allen when the Bridgewater, N.J.–based drug company Aventis was considering tests of whether compounds that block interleukin-5 work as treatments for asthma attacks. When exposed to virtual allergens and no drugs, Bill’s asthma attacks lasted 3 days, much longer than is typical for people with asthma.
Simulated injections of interleukin-5 blockers prevented Bill’s asthma attacks but not Allen’s.
Because Bill’s asthma didn’t seem to reflect real life and Allen didn’t respond to the interleukin-5 blockers, Aventis didn’t pursue these compounds as potential asthma therapies. The Entelos model seems to have been accurate. Despite promising animal studies, when other companies recently tested interleukin-5 blockers in people, they found that the compounds have much less effect than the researchers had originally expected.
Each simulation of a disease begins by modeling the normal physiology and interaction of the organs involved. “We are striving for a whole-body approach to health and disease,” says Jeff Trimmer of Entelos. “We want to use [our models] to understand how a person gets sick.” Even when models don’t seem to simulate what happens in real life–as in Bill–the findings can help researchers better understand physiological factors that are important in causing diseases, says Trimmer.
To date, computer simulations of basic human biology are closely intertwined with the commercial quest to develop new drugs quickly and inexpensively. Pharmaceutical companies can now take more than a decade to move a compound from the lab to the clinic, and many potential drugs fall by the wayside in that journey. Companies report that about 20 percent of the drugs that succeed in preliminary trials are abandoned once doctors test them in large, expensive clinical trials: Either they don’t work as expected, or troubling side effects emerge.
The current system is so bad that anything would be an improvement, says Lewis Sheiner of the University of California, San Francisco, who models drug responses.
Drug companies hope that computer simulations will focus their efforts on the candidate drugs most likely to work in people. With biological models of cells, organs, and metabolic pathways, industry scientists can pick out genes or proteins involved in a disease and then sort through various drug candidates. Only the most promising ones would then be tested in cells and animals. Such modeling is gaining proponents, says Karin Jorga of Hoffmann-La Roche in Basel, Switzerland.
Computer simulations may also reduce the number and size of human trials that a company has to conduct on the way to a drug’s approval by the Food and Drug Administration, Jorga says. The simulations can indicate the most efficient trial designs to reveal beneficial effects. Some computer models that simulate how drugs are broken down in the body can predict the most effective dose, she says. At a meeting on drug effects in the Netherlands earlier this year, Jorga reported that such modeling reduced the time Hoffmann-La Roche needed to develop and test a new formulation of a drug for preventing bone loss. With the new formulation, people could take the drug once a month rather than once a day.
Another example of how computer simulations have guided drug design is a virtual heart developed by Denis Noble of the University of Oxford in England. The model consists of a mass of virtual cells, each processing virtual sugar and oxygen. On a computer screen, this heart beats just like the real thing. The virtual heart can be programmed to develop different diseases, and scientists can see how it responds to treatment with different drugs.
Noble’s model has been used, for example, to predict whether one drug is likely to cause abnormal heart rhythms, a widely recognized, potentially dangerous side effect. Computer simulations, however, aren’t good at predicting unexpected side effects.
Greater advantages could result if biomedical researchers combine the models they’re developing. Entelos is trying to hook together models of several different diseases to increase its chances of detecting side effects or spotting dangerous drug interactions. For example, heart disease, obesity, and diabetes tend to occur in the same patients, Trimmer says, so it would be useful to model all those ailments at the same time.
Bassingthwaighte and other researchers are combining individual computer models of genes, proteins, cells, organs, and metabolic pathways. With such a set of coordinated models, he suggests, medical scientists might see effects of disease that are hidden by the complexity of the human body.
This effort–sometimes called the virtual human or physiome project–is just beginning. Many technical and scientific hurdles remain: getting the models to communicate with one another, finding the powerful computers to run such simulations, even gathering the data needed to develop some of the models.
And not everyone is convinced that unified models will offer helpful information. “If you’re an engineer, it isn’t useful to use a quantum mechanical model of the world to talk about girders and bridges,” says Sheiner. “Getting the scale of the model to match the scale of the question is tricky, so that working from the genes up may not be an efficient way to determine the best ways of treating particular diseases.”
Modeling medical care
Some researchers are applying computer simulations to an even broader realm of health care. Rather than studying cells and organs, researchers associated with the Oakland, Calif.–based health-care plan Kaiser Permanente have created a computer simulation of a virtual world where people develop diseases–asthma, diabetes, and heart disease so far–and go to doctors for tests and treatments. Called the Archimedes model, it may help Kaiser Permanente set guidelines for patient care and find ways to monitor caregivers’ performance, says physician David M. Eddy, a consultant to Kaiser Permanente. He and physicist Leonard Schlessinger of Kaiser developed the model.
Archimedes may also predict results from clinical trials that simply can’t be carried out in the real world. It may also give physicians results more quickly than would be possible with an actual trial. Eddy cites a variety of obstacles for carrying out real-world trials: “the pace of innovation, the high cost of doing research, the long follow-up times required, the large number of options to be compared . . . and the unwillingness of the world to stand still until the research is done.”
All those factors “severely limit our ability to evaluate all the options [for patient care] through clinical research alone,” he adds.
The model takes into account many influences on disease in real people, says Eddy. For example, each virtual patient has a virtual liver and pancreas. The health of these organs affects the concentration of sugar in a virtual patient’s blood. When sugar concentrations rise high enough, a patient may experience symptoms of diabetes, such as thirst or frequent urination, and go to a doctor. Although the underlying causes of diabetes aren’t well understood, Archimedes uses equations that reproduce the disease’s known characteristics, such as the observed incidence of the disease in different ethnic groups.
“As in reality, each patient is different,” says Eddy. Some patients may take medicine or follow a doctor’s recommendations more meticulously than others do. Moreover, patients may receive care of different quality. Reflecting studies of doctors’ habits, a virtual primary-care physician is less likely to accurately follow treatment guidelines than is a virtual specialist. These factors, in turn, affect a number of physiological variables in the computer, as in life.
For example, different simulated diet and exercise plans reduce the cholesterol, blood pressure, and weight of the virtual patients by different amounts. Those changes then affect the likelihood that, say, an overweight patient would develop diabetes or heart disease.
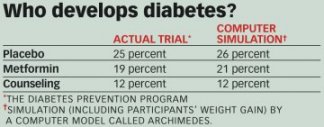
At the annual meeting of the American Diabetes Association in June, Eddy reported results from an Archimedes simulation of a real-life, large clinical trial. The actual trial was known as the Diabetes Prevention Program (SN: 9/8/01, p. 159: Available to subscribers at Walking and eating for better health). Mimicking that trial, the virtual one enrolled virtual overweight people with a variety of risk factors for developing diabetes. It randomly assigned them to receive intensive counseling about diet and exercise, a drug called metformin that increases a person’s sensitivity to insulin, or a placebo.
To program the model, Eddy and his colleagues used results from a variety of real-life studies about the effects of various interventions on weight and a variety of other risk factors for diabetes. The first time Archimedes simulated the actual diabetes trial, the virtual results were very close to the real results–as close as one would expect results of two actual trials to be, says Eddy. The simulation almost exactly mimicked the clinical trial when the modelers included the weight regained by participants.
“If we could not have afforded that trial, we could have predicted the results quite well using this model,” says Richard Kahn of the American Diabetes Association in Alexandria, Va. “The [simulated] health-care system responded very much like the real one, and that suggests that Archimedes could help improve the way we treat patients.”
Simulating reality
Despite the promise of technology, “no model is perfect,” cautions Bassingthwaighte. “The limitation is our database of knowledge. If we don’t understand it, we can’t model it.”
Thanks to the Human Genome Project, which recently decoded essentially the entire human genome, researchers now have a large database of genes and proteins, he says. A new field of research, called proteomics, is adding information about the expression of proteins and their interactions in cells. However, Bassingthwaighte cautions, relatively little is known about how most of those molecules affect cells and organs.
He expects that extensive experimenting will be required to tease out that information.
Likewise, simulations of disease processes and health care can reproduce what’s known, but can’t always identify an unexpected side effect. For instance, even today, a computerized model probably wouldn’t predict the damage that the diet drug fenfluramine causes to heart valves, as revealed in studies in 1997 (SN: 10/18/97, p. 252: http://www.sciencenews.org/sn_arc97/10_18_97/bob2.htm), says Trimmer. It’s for such reasons that he notes, “This [simulation] technology is not a replacement for traditional research but a complement to it.”
“The fundamental idea is that if you can model a real thing on a computer, you can answer a lot of questions without experimenting with the real thing,” says Sheiner. “While models are far from perfect because they depend on imperfect information, they can give you an intelligent basis for making decisions when you’re faced with uncertainty.”
Early progress with computerized biomedicine has modelers enthusiastic about its future. “My goal is to move medicine onto a quantitative footing,” Eddy says.
“There’s still a long way to go, but simulations will certainly help. We’ve shown that this plane will fly.”
****************
If you have a comment on this article that you would like considered for publication in Science News, please send it to editors@sciencenews.org.
To subscribe to Science News (print), go to https://www.kable.com/pub/scnw/subServices.asp.
To sign up for the free weekly e-LETTER from Science News, go to http://www.sciencenews.org/subscribe_form.asp.