In what some scientists are describing as the most sweeping innovation in gene sequencing in the past 25 years, researchers have developed a tool that can read out DNA’s genetic letters up to 100 times as fast as the standard technique does.
“It has the potential to really change the way we do things with genomes,” says H. Chad Nusbaum, codirector of genome sequencing and analysis at the Broad Institute, a genetic research center jointly operated in Cambridge, Mass., by Harvard University and the Massachusetts Institute of Technology.
The new approach, developed by 454 Life Sciences Corp. in Branford, Conn., sidesteps a bottleneck in conventional gene sequencing. In machines employing that scheme, known as the Sanger method, only 96 stretches of DNA can be sequenced at any one time. Jonathan M. Rothberg of 454 Life Sciences and his colleagues have now simultaneously sequenced hundreds of thousands of snippets of DNA. The work is described in an upcoming Nature.
With either approach, technicians first chop DNA into segments. Sanger sequencing next puts the segments in bacteria for replication. Removed from the bacteria, those copies then undergo more replication and fluorescent tagging. Finally, they are fed into a 96-tube device that reveals the sequence of the individual base pairs that make up the DNA segments.
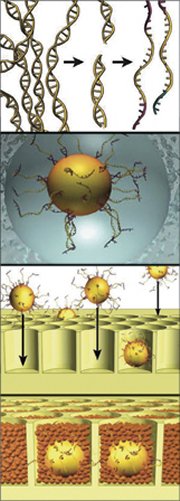
Dispensing with bacteria altogether, the 454 Life Sciences approach uses a test-tube process to coat each of hundreds of thousands of microscopic beads with DNA segments. Each bead carries millions of copies of one segment. Next, each bead is deposited in its own tiny well in a credit card–size plate monitored by a dense photoreceptor array that feeds data to a computer. As chemical reactions occur in each well, they yield flashes of light signifying which base pairs are located along the segment.
The Sanger method routinely sequences DNA segments containing 800 base pairs, while the 454 Life Sciences report describes sequencing only 100-base-pair segments. Because it’s far more difficult to puzzle out the complete genome from the shorter segments, the new approach has limited usefulness for analyzing large, repetitive genomes, such as those of mammals, says Jane Rogers, head of genome sequencing at the Wellcome Trust Sanger Institute in Hinxton, England.
Rothberg told Science News that his team is modifying the method so that it will work with longer strands. The team has already sequenced segments of 500 bases, he says.
Although the 454 Life Sciences machine costs half-a-million dollars—about $150,000 more than conventional equipment—the higher throughput drives down the cost of sequencing, says Rothberg.
Lowering sequencing costs is a major goal of geneticists. Current technology can spell out a person’s genome, or full genetic code, at a cost in excess of $10 million, according to Geoff Spencer of the National Human Genome Institute in Bethesda, Md. Cutting that price drastically could make it more feasible for doctors to determine an individual’s propensity for certain diseases.
Although Rothberg and his colleagues are the first to bring to market a contender to the Sanger method, many other groups are developing challengers. In an upcoming Science, for instance, Jay Shendure of Harvard Medical School in Boston and his colleagues report a low-cost sequencing machine that they built on a budget of $140,000.
“The new-sequencing-technology game is a horse race,” Nusbaum says. “454 is way ahead right now,” he says, but no one yet knows who the ultimate winner or winners will be.





