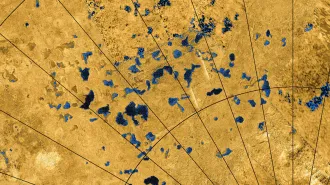
A TINY WORLD Eric Betzig (left), Stefan Hell (center) and W.E. Moerner share the 2014 Nobel Prize in chemistry for developing high-resolution microscopy techniques.
From left: Matt Staley; © Max Planck Institute for Biophysical Chemistry; Linda A. Cicero/Stanford Univ. News Service
Breaking the limits of light microscopy by using fluorescence has won three scientists the 2014 Nobel Prize in chemistry. Eric Betzig, a physicist at the Howard Hughes Medical Institute’s Janelia Research Campus in Ashburn, Va., Stefan Hell, a physicist at the Max Planck Institute for Biophysical Chemistry in Göttingen, Germany, and W.E. Moerner, a physical chemist at Stanford University, have been awarded the prize.
Using microscopy techniques that the prize winners developed, researchers can now peer into the depths of bacterial cells, watch neurons shift shapes in learning brains and glimpse clumped-together proteins in diseases such as Alzheimer’s, Huntington’s and Parkinson’s (SN: 6/15/13, p. 20).
“It has provided a window into the cell,” says Catherine Lewis, a biophysicist at the National Institute of General Medical Sciences in Bethesda, Md.
If traditional light microscopy were likened to looking at an anthill, fluorescence microscopy would be like seeing individual ants — but better, said chairman of the Nobel Committee for Chemistry Sven Lidin, an inorganic chemist at Lund University in Sweden. “You can look at the legs of the ants,” he said, and even see bumps and bruises.
A human hair’s width is about 500 times this limit, Lidin said at the press conference announcing the prize, as he plucked a strand from his own head. So objects of that size can be easily studied with traditional light microscopy. But most of the machinery of living cells is orders of magnitude smaller.
Hell and colleagues found that, using traditional microscopes, cell parts smaller than 200 nanometers across look like blobs. But he wanted to see within the blobs.
“The scientific community wasn’t very receptive to the idea of overcoming the diffraction barrier,” Hell said at the press conference. People believed “that doing something about it was — pardon me — kind of crazy.”
He added, “Eventually I realized there must be a way by playing with the molecules.”
In 2000, Hell and colleagues shot a combination of colored lasers at fluorescent molecules. The first laser lit up a wide group of molecules while a second laser with a donut-shaped beam knocked out the glow of any molecules in its path. This left a tiny circle, smaller than 200 nanometers across, spotlighted in the center for scientists to observe.
Though researchers had devised another method of spying on such wee molecules, that technique, called electron microscopy, requires cells to be killed, sliced, stained with chemicals and blasted with intense radiation, Lidin said. With fluorescence microscopy, researchers can observe bacteria and other cells without destroying them, he said. “They can be studied in real time while they live and prosper.”
To watch the happenings inside cells, Moerner and Betzig working separately each used the same microscopy trick as Hell but with a twist. Instead of blocking light from a swath of molecules, the researchers engineered light switches for molecules, explains microscopist Brian English, Betzig’s colleague at Janelia. In 1997, Moerner and colleagues reported that zapping fluorescent molecules with different wavelengths could cause individual molecules to light up or black out.
Building on that finding, in 2006, Betzig and colleagues used similar molecules to view a lone membrane protein from a mammalian cell. The method, he says, could one day “tell us how inanimate molecules come together to create animate life.”
Chemist Richard Zare of Stanford marvels at the advances. “We all in chemistry — we believed in molecules,” he says. “But no one had ever seen one” before these techniques, he says.
The three winners will divide equally the roughly $1.1 million prize.