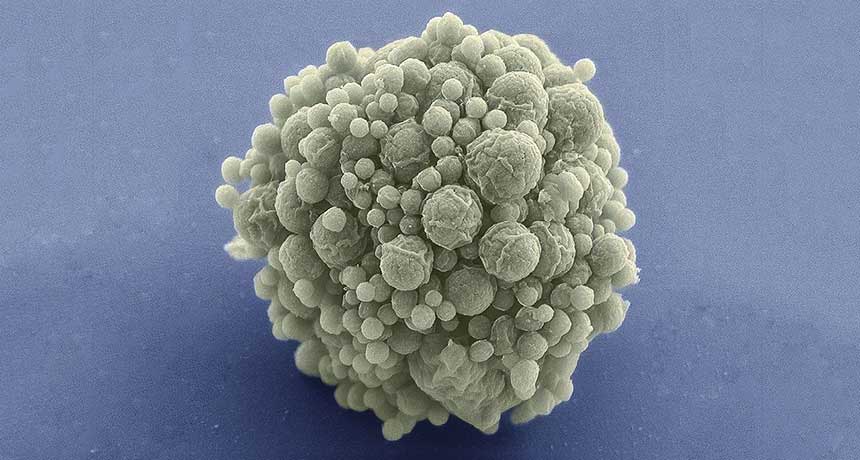
MOST MINUSCULE Named syn3.0, engineered bacteria developed at the J. Craig Venter Institute can survive and function with just 473 genes.
Mark Ellisman/National Center for Imaging and Microscopy Research
![]() One of biology’s biggest achievements of 2016 was intentionally as small as possible: building a bacterium with only 473 genes. That pint-size genetic blueprint, the smallest for any known free-living cell, is a milestone in a decades-long effort to create an organism containing just the bare essentials necessary to exist and reproduce. Such “minimal genome” cells might eventually serve as templates for lab-made organisms that pump out medicines, make innovative chemicals for industry and agriculture, or churn out other molecules not yet imagined. The project also identified genes crucial for the microbe’s survival yet largely unfamiliar to science, highlighting major gaps in researchers’ grasp of life’s playbook.
One of biology’s biggest achievements of 2016 was intentionally as small as possible: building a bacterium with only 473 genes. That pint-size genetic blueprint, the smallest for any known free-living cell, is a milestone in a decades-long effort to create an organism containing just the bare essentials necessary to exist and reproduce. Such “minimal genome” cells might eventually serve as templates for lab-made organisms that pump out medicines, make innovative chemicals for industry and agriculture, or churn out other molecules not yet imagined. The project also identified genes crucial for the microbe’s survival yet largely unfamiliar to science, highlighting major gaps in researchers’ grasp of life’s playbook.
The newly engineered bacterium was praised as a technical triumph. In 2010, researchers at the J. Craig Venter Institute in La Jolla, Calif., had stitched together a copy of the entire genome of the bacterium Mycoplasma mycoides and popped it into the cell of another bacterium whose genome had been removed. But that “synthetic cell,” dubbed JCVI-syn1.0, contained a full copy of an existing genome. With more than 1 million chemical building blocks of DNA, including 901 genes, it was far from minimal.
The latest version, JCVI-syn3.0, reported in March in Science (SN: 4/16/16, p. 6), has roughly half that much DNA. It’s also the first cell built using human design principles: One segment of the genome has genes for various processes, such as DNA repair, grouped together rather than scattered willy-nilly. Abandoning the untidiness of evolution for a logic-driven blueprint enables a “plug and play” approach, says Daniel Gibson, a member of the JCVI team. To tinker with a metabolic process such as glycolysis, for example, “Rather than changing one gene, then another, then another, you could pop out a whole module and then pop in a new one.”
Making such fundamental changes to the genome while still getting a functioning cell is noteworthy, says genome scientist George Church of Harvard University. “They could have found that, no matter what they did putting it together, it broke,” Church says.
The potential of synthetic cells is enormous, says Claudia Vickers, a biotechnologist at the Australian Institute for Bioengineering and Nanotechnology in Brisbane. Scientists have succeeded in engineering existing organisms such as yeast to help make, for example, malaria drugs. Now little cellular factories designed to be highly efficient and tailored to specific tasks are within sight, Vickers says.
Story continues below graph
Only the essentials
This year scientists introduced a synthetic cell with just 473 genes — the smallest known genetic blueprint for any free-living cell. Called JCVI-syn3.0, it is built on a previous effort dubbed JCVI-syn1.0, which had 901 genes. At 525 genes, the slow-growing Mycoplasma genitalium also has a tiny genome, especially compared with the illness-causing Haemophilus influenzae and Escherichia coli.
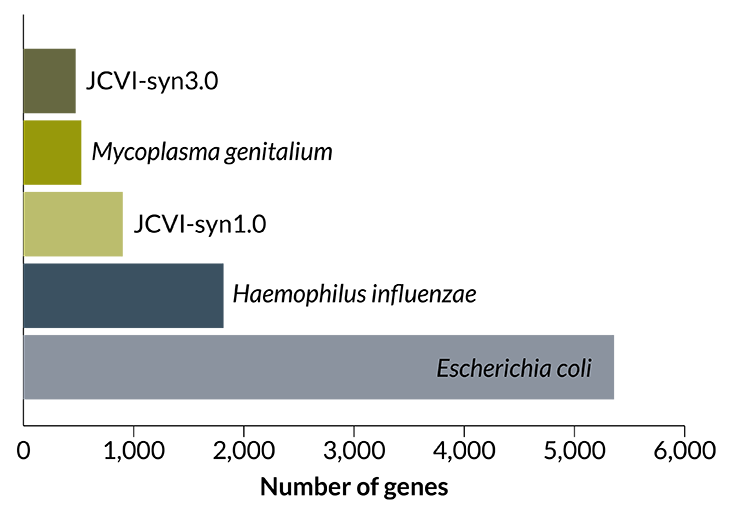
Sources: E. coli: Genome/NCBI/National Library of Medicine; all others: C.A. Hutchison et al/Science 2016
The techniques used to build JCVI-syn3.0, especially when considered alongside other engineering tools such as the recently developed CRISPR/Cas9 system (SN: 9/3/16, p. 22), are a meaningful step toward the once-distant goal of self-replicating minimachines. “It’s important for the future it allows us to imagine,” Vickers says.
Since announcing JCVI-syn3.0, the team has used the same engineering techniques to turn the fast-growing bacterium Vibrio natriegens into a laboratory workhorse. The engineered Vibrio — dubbed Vmax — cuts the time it takes to do particular lab experiments in half compared with the original, Gibson says.
The minimal genome effort also aims at a larger philosophical question: What is life? In a lecture in 1984, origin-of-life expert Harold Morowitz discussed how studying the small and simple Mycoplasma genome might invigorate basic biology in much the way that studying the hydrogen atom sharpened questions for physics and chemistry. (Morowitz died in March, two days before the JCVI-syn3.0 work was published online.)
Many scientists, for example, were stunned to learn that JCVI-syn3.0 had 65 genes with no known function that were nevertheless required for survival. “This is one of our best studied organisms, and we haven’t the foggiest idea what those genes are doing,” says evolutionary genomics expert Laurence Hurst of the University of Bath in England. “It’s a brilliant result.”