Big data studies come with replication challenges
The truth can be hard to find with millions of data points and lots of room for error

FOOL'S GOLD As researchers pan for nuggets of truth in big data studies, how do they know they haven’t discovered fool’s gold?
James Provost
In my quest to explore the unknown frontier inside my own body, I stumbled upon one of the most intractable problems facing science.
The issue, irreplicable results, is a dark cloud looming over the life and social sciences. My encounter began when I sent identical stool samples to two microbiome sequencing services (SN Online: 6/17/14). The microbiome is the community of microbes that live in and on the human body, and studies had suggested that it shapes health and even behavior. My goal was to find out what bacteria inhabit my intestines, the most microbe-packed part of the body.
I thought the process would be straightforward: the two services, American Gut and µBiome, would examine the DNA from the microbes in my gut and tell me what was in there. But when the results came back I was no wiser than before — just confused. The profiles presented by the services showed wildly different results. For example, they reported almost completely opposite readings on the proportions of Firmicutes and Bacteroidetes, two of the major phyla of bacteria found in the human gut. This was frustrating because the mix of these two may determine whether someone is obese or not and affect other aspects of health. I thought that, at a minimum, I would learn how many of these major players inhabited my gut (
).
Although the bacteria, viruses and fungi that microbiome scientists study are microscopic, the amount of information needed to catalog the microorganisms and figure out their effect on the body is massive. For one sample like mine, there may be hundreds of different types of bacteria, and thousands of bits of DNA to sequence and analyze. To tell me how my personal mix stacks up, the researchers compared my sample with thousands of other people’s. But to discover the impact of different microbes on health, scientists might analyze hundreds or thousands of samples, each containing its own ecosystem of myriad microbes. If two labs couldn’t get my sample right, what does that say for the vast studies cataloging the bacteria involved in human health and disease?
When I tweeted and blogged about the anomalous findings, people who study the microbiome confessed that they were not surprised my results differed so starkly. They had encountered this problem before, and now they were preparing to tackle it. I was invited to watch.
Microbiome researchers aren’t the only ones having a tough time replicating results. Retractions and corrections in the scientific literature are on the rise. Scientists are hotly debating both the sources of and solutions to the problem that is rocking science (SN: 1/24/15, p. 20).
It is hard enough to duplicate findings from studies with a handful of mice or people. But extend the work to include thousands or millions of data points collected from huge numbers of research subjects — the kind of work done in the expanding field of genomics, for example — and the room for error grows by leaps and bounds.
Microbiome research is just one of many flavors of the “big data” projects that have become ubiquitous in the life sciences. For genome-wide association studies, researchers track hundreds of thousands of DNA variants in tens of thousands of people to find genetic contributions to common diseases. In sequencing studies, geneticists are collecting billions of bits of DNA data on hundreds to thousands of research subjects from lab animals to humans. Brain scientists are attempting to map all of the 86 billion neurons in the human brain and catalog the trillions of connections they make with other neurons. The list goes on.
Big data projects are officially defined as those that generate so many pieces of information that computers are needed to sort through it all. But that doesn’t begin to capture the scope of these efforts.
Daniel MacArthur, a geneticist at the Broad Institute of MIT and Harvard, formed a coalition called the Exome Aggregation Consortium with 23 other scientists and their research associates. The group has pooled genetic data from the exomes, or protein-coding parts, of more than 90,000 people’s genomes. The database holds about 925 terabytes of raw data — more than nine times the size of the print collection of the Library of Congress. And more genomes are being added all the time. Plenty of other researchers are generating their own enormous masses of data.
Buried in that data are potential gold mines. A recent study of 2,430 bacterial genomes, for example, showed that friendly microbes can make 44,000 small molecules, including some that could be useful antibiotics (SN: 10/18/14, p. 8).
Researchers are sifting through mounds of data to find and scrutinize similar nuggets to develop better drugs or make connections between genetic variants and diseases. Yet, as science moves toward big data endeavors, so grows the concern that much of what is discovered is fool’s gold.
Just keeping track of big data is a monumental undertaking. Sharing the data with other researchers, a critical piece of transparency and efficiency in science, has its own set of problems. And the tools used to analyze complex datasets are just as important as the data themselves. Each time a scientist chooses one computer program over another or decides to investigate one variable rather than a different one, the decision can lead to very different conclusions.
For instance, two groups of researchers applied different analyses to one dataset containing gene activity measurements from mice and humans injured by trauma, such as burns or blood infections. One group concluded that mice are terrible stand-ins for people with inflammation caused by trauma (SN: 3/23/13, p. 10). The other group decided that the rodents are excellent human analogs (SN: 9/20/14, p. 14). Same data, opposite results.
Optimists within the scientific community hope to avoid at least some pitfalls by learning from others who have conquered similar challenges before. For example, researchers who study gene activity with devices known as microarrays, available since the mid-1990s, were among the first biologists to encounter the big data dilemmas. They have stepped through technical problems and are perfecting ways to allow disparate research groups to directly compare their data.
Sorting out the weaknesses
Studies of the microbiome produce some of the hottest papers in biology today. But, as I discovered, results in one lab don’t always match up with those from another. For Rashmi Sinha, an epidemiologist at the National Cancer Institute, and others, the disagreement between labs means that conclusions about how microbes affect health can’t be fully trusted. With nearly every aspect of human biology dependent on microbial actions, microbiome researchers need to be able to count on their data.
The first step in slaying any dragon is learning its weaknesses. Sinha masterminded a plan to probe for soft spots in the way scientists collect, process and analyze micro-biome data. The project is known as the MBQC, for microbiome quality control. Many of the top labs in the field eagerly signed up.
Last autumn, about 60 microbiome researchers and observers met in Rockville, Md., to talk about testing for vulnerabilities in microbiome studies.
Sinha and microbiologist Emma Allen-Vercoe of the University of Guelph in Canada had prepared 96 standardized samples of bacteria or DNA for the researchers to examine. In the project’s pilot phase, 15 laboratories handled and sequenced the microbiomes, then handed their data to nine labs for computer analysis.
Each lab was encouraged to follow its normal procedures and closely document each step. “It’s surprising how many little things each lab chose to do differently,” says Curtis Huttenhower, a computational biologist at the Harvard School of Public Health. For instance, researchers used diverse methods to crack open the bacteria and pull out the DNA. Analysis methods varied widely as well.
The idea wasn’t to judge whose choices were better. Instead, the researchers wanted to know which steps injected chaos into the system.
A few procedural decisions affected the final results. One lab that studies the vaginal microbiome used a unique set of tools, called PCR primers, to make copies of the DNA. That lab counted a different amount of diversity of microbes than the rest of the labs. The DNA extraction methods mattered, too, as did the analytical techniques.
That’s the boring news, Huttenhower says. True, such variables are a source of error that could lead researchers to detect patterns that aren’t due to the underlying biology. But the exciting result, he says, was that even with the technical differences, researchers were still able to reliably differentiate samples from sick people from those of well people. The result gives him hope that veins of biologically meaningful information run through the mountains of data.
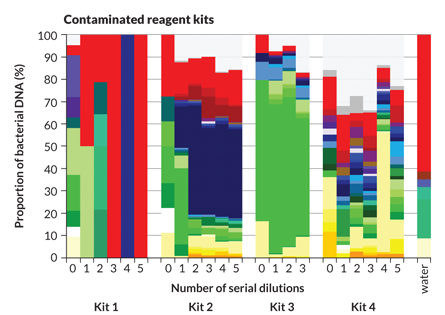
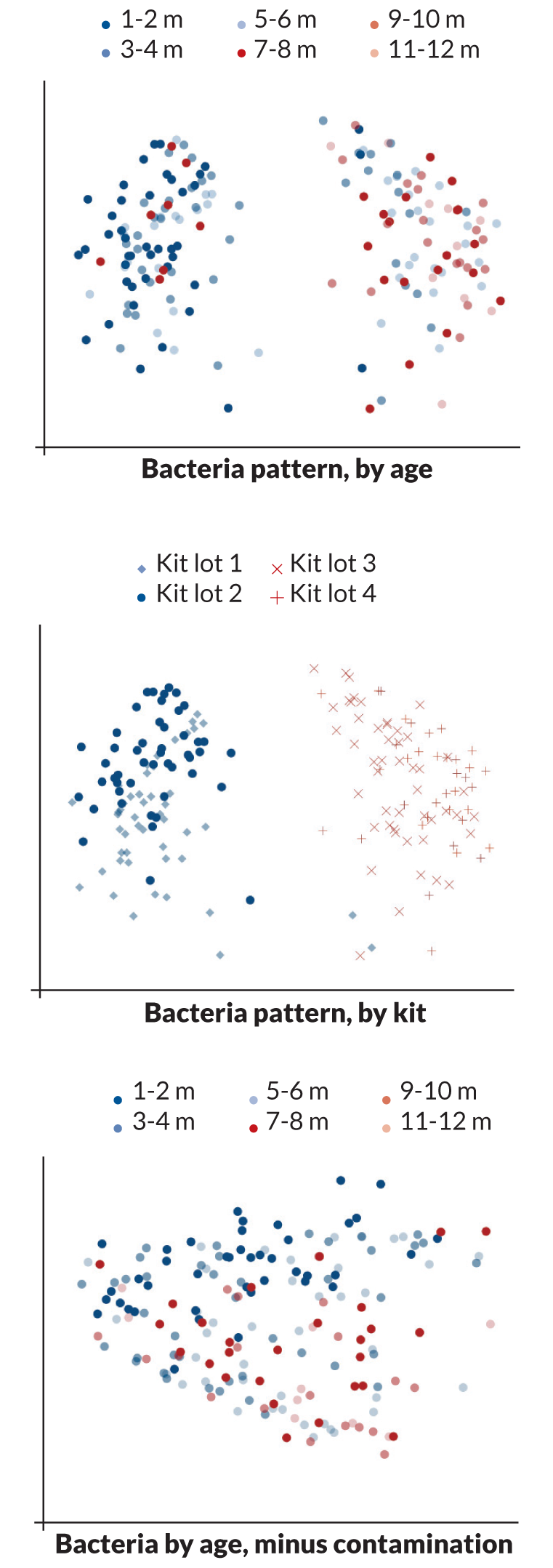
Sometimes, however, hidden menaces are so noisy that they drown out the real biological message. Contamination is one such menace, Susannah Salter of the Wellcome Trust Sanger Institute in Hinxton, England, and colleagues discovered. Sterile water and reagent kits that scientists use to pull DNA from microbial samples may already contain significant amounts of bacterial DNA, the researchers reported last November in BMC Biology. Contaminating DNA may dominate samples, throwing off results.
Salter and her colleagues described one such instance: A study of how children’s microbiomes develop that took place in a refugee camp on the border between Thailand and Myanmar. A group of infants born in 2007 and 2008 had their noses and throats swabbed every month until they were 2 years old.
At first it looked as if soil bacteria such as Achromobacter and Herbaspirillum are the first to grow in infants’ noses and mouths. But the finding made Salter suspicious. “The things we were seeing were not normal human bugs,” she says.
Because her Thai colleagues kept meticulous records, Salter was able to determine that the soil bacteria were not growing in babies’ noses. Instead, DNA from soil bacteria had somehow contaminated two of the kits used to process the samples, completely obscuring the organisms that were really present.
Contamination doesn’t end with lab reagents. It can be enshrined in digital format in large databases, Steven Salzberg of Johns Hopkins University and colleagues reported in November in PeerJ. The researchers found that the genome of Neisseria gonorrhoeae, the organism that causes gonorrhea in people, was contaminated with stretches of DNA that actually came from cows and sheep. Four other genomes the researchers randomly selected from the public database GenBank also contained sequences from other species, indicating that many more genome records may also be tainted.
These types of challenges are to be expected when working with big data, researchers say.
“Anything that provides a lot of very sensitive data provides a lot of truth and a lot of noise,” Huttenhower says. Scientists need to know about and account for the noisemakers in their studies. Huttenhower prefers telling people what to look out for rather than being prescriptive. Forcing scientists to conform to a single protocol would be a mistake, he says.
Go big or go home
While it is easy to get lost in big data and see patterns where none exist, sometimes the problem with big data is that it’s not big enough.
MacArthur and other scientists in the Exome Aggregation Consortium are trying to track down very rare mutations that cause diseases. MacArthur focuses on muscle diseases, such as muscular dystrophy and congenital myopathy.
He and colleagues must comb through about 30 million DNA bases that make up one person’s exome to find the one or two mutations that cause the disease. That task would be hard enough, but it is further complicated because even a healthy person’s genome carries about 20,000 to 30,000 genetic variants. How can scientists tell whether what they’ve found is really a disease-causing mutation and not a benign rare variation? Often they can’t.
A study published in Science Translational Medicine in 2011 found that 27 percent of variants identified as the causes of inherited rare diseases either turned out to be fairly common or were mis-labeled. To MacArthur, the implications are clear: “All of us who have done rare disease discovery in the last decade have almost certainly misdiagnosed patients.”
The solution to the problem? Go bigger. MacArthur and colleagues realized that pooling data from huge numbers of people in the Exome Aggregation Consortium would give them a better picture of just how common variants are in the population. Armed with that knowledge, researchers can be more confident that the mutations they discover really are rare and the likely cause of a disease.
“The impact of big data on science is unquestionably a force for good,” says MacArthur. “It sweeps away false positives.”
But creating the database was no easy task. Each project that contributed data generated it differently, MacArthur says. He and others spent nearly two years developing software to harmonize the data from disparate sources.
Messy data
Big data sharing is an ordeal that is all too familiar to Santiago Pujol, a civil engineer at Purdue University in West Lafayette, Ind. Pujol and colleagues are creating a virtual platform to allow 15 labs involved in an earthquake engineering project sponsored by the National Science Foundation to store and share their data.
“What we got was quite challenging,” Pujol says. Sometimes data files would arrive with no explanation of the type of information contained within, or measurements would be labeled but didn’t include units. “You didn’t know if they were measuring in millimeters or inches,” Pujol says. In short, it was a mess.
Pujol’s experience is not unusual. For the MBQC, Huttenhower thought he had given the microbiome researchers precise instructions for how to present sequence data to the analytical labs, but he says he still got a variety of file types and data formats that had to be reconciled before the computer programs could analyze them.
Pujol and others say that standards for presenting and storing data could go a long way toward making research of all types more reliable.
Safer ground
Big data researchers are hoping to learn from the microarray pioneers who have already tackled some of these replicability challenges. Researchers who use these devices to measure gene activity were some of the first prospectors in life science’s big data rush.
Microarrays use a red and a green dye to measure gene activity and are widely used in studies comparing how disease or environmental conditions, such as exposure to chemicals, affect cells.
Kristopher Kilian, a chemist at the University of Illinois Urbana-Champaign, worked at microarray-maker Rosetta Inpharmatics in the early 2000s when the technology was catching fire. The company was fielding complaints from users that results of experiments changed with the seasons. Then some of the microarrays caught “measles,” the company nickname for a strange pattern in which red dots ringed in green appeared on the microarrays.
The measles struck when the company built its new microarray processing facility next to a freeway, and they peaked with rush hour traffic. Finally, Kilian’s group determined that ozone produced by cars was degrading the red dye, and suggested in Analytical Chemistry in 2003 that researchers keep levels of the gas low. Ozone also varies by season, with higher levels floating around in the summer, and could explain why researchers were getting different results at different times of year.
Carbon filters that scrub ozone out of the air dramatically improve the reliability of microarray experiments, another group of researchers
in
BMC Biotechnology
.
Working out technical problems was just the first step. It took several years of software development to make experiments done in various labs using the same type of microarray comparable with each other. And researchers still have trouble reconciling experiments that rely on microarrays produced by different companies, and yet more difficulty comparing the results with new technologies such as RNA sequencing, a more sensitive way to measure gene activity. Researchers adopting the new technology would like to reconcile their fresh-off-the-sequencer results with older microarray findings, but currently they can’t.
Even that hurdle may be surmountable. Biomedical engineer Sarah Munro of the National Institute of Standards and Technology in Stanford, Calif., and colleagues have developed a set of 96 standardized RNAs for use as internal quality controls to tell how well researchers are performing each step of their experiment. The standards should allow researchers to calibrate their results to those from other labs and possibly match up microarray data with RNA sequencing results. Computer software that the researchers call a dashboard allows scientists to try out several types of analysis on their data to see how the final outcome might change.
Eleven of 12 labs that tried the standards showed consistent performance, Munro and colleagues reported last September in Nature Communications.
“You spend all your time doing these experiments, so you want to know you’re getting it right,” says Munro. She hopes the standards will help other researchers better evaluate and replicate results. “It’s about people being able to communicate their measurements and have confidence in them,” she says.
It has taken more than 15 years for microarray technology to develop enough so that scientists can easily compare their data. Researchers conducting other types of big data studies hope that the lessons learned from more mature fields will help catapult them past pitfalls to safer ground where data can be trusted. With data shooting from sequencing machines and other high-throughput laboratory equipment like water from a fire hose, it has never been more important that researchers learn what it takes to make their results as reliable as possible.
This article appeared in the February 7, 2015 issue of Science News with the headline “Big Data, Big Challenges.”