Fly over any baseball stadium when the home-team batter slams a double in the gap with two men on base, and you’ll see a crowd of fans rising in unison, arms waving wildly in the air. You’d think you were viewing typical baseball fan behavior.

Witness this scene at ground level, though, and you’ll get a different picture. While a majority of fans participate in the cheering, others are sipping beer, attending to scorecards or roaming the walkways in search of a hot dog.
Such displays of individuality shouldn’t come as a surprise — people often react differently to the same circumstances, after all. But life’s individuality thrives at much smaller scales than the human body.
Cells, for example, are not all alike. Clusters of single cells, even of all the same type, show cell-to-cell differences in appearance, growth and behavior. This holds true for single-celled organisms too. Individuals within a colony of identical bacteria, for instance, will behave in different ways, even under the same conditions.
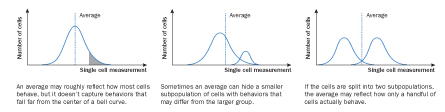
For decades, scientists were stuck with an aerial view of cell behavior. Experiments carried out on large pools of cells obscured information on the activities and makeup of the individuals. Difficulties in tracking molecules inside a single cell forced researchers to content themselves with averages derived from whole populations.
Now scientists are finding ways to zero in. Improvements in imaging techniques make it possible to view tiny, transparent structures and molecules within cells that are otherwise nearly invisible. New ways of separating cells during study enable scientists to sort through thousands at once, distinguishing subpopulations and studying multiple traits in just one go.
“We used to think that we could take a million cells and grind them up and make measurements on those million cells,” says Nancy Allbritton, a biomedical engineer at the University of North Carolina at Chapel Hill. “And whatever measurements we made on those million cells would reflect what’s going on in a single cell.”
Scientists today know that averaging the constituents of a million cells often gives no clue to what’s going on at the cellular level. Cells thought to be alike have been found to produce different numbers and types of proteins. Beyond providing interesting insights into basic biology, understanding this individuality may have medical consequences — illuminating characteristics that can make a cell more amenable to a particular drug or more vulnerable to an allergen or viral infection.
Though in many ways single-cell analysis is still in its infancy, the field is quickly expanding. Researchers are working to improve technologies and to figure out how to make sense of the data.
Lighting the way
Of the millions of different living things in the world — large and small, plants, animals and microbes — all have in common the fact that they are made of one or more cells. Scientists have been observing cells since the mid-17th century, when Dutch scientist Antony van Leeuwenhoek discovered bacteria and, later, blood cells. Through the years, microscopes and staining methods have allowed researchers to view and describe the basic structures and internal workings of various cell types — understanding how skin cells, for example, differ from cells in the heart or kidneys.
Distinguishing unique features among cells of the same type has proven to be more difficult, because cells are so small. A typical human cell is about one one-hundredth of a millimeter across. Microbes are even smaller. Conventional light microscopes are constrained by a physical law demanding that the size of an object being imaged be no smaller than about half the wavelength of the light used to produce that image. To see anything as small as a molecule within a cell (the scale at which much of the work gets done), some tool other than traditional light microscopy is needed.
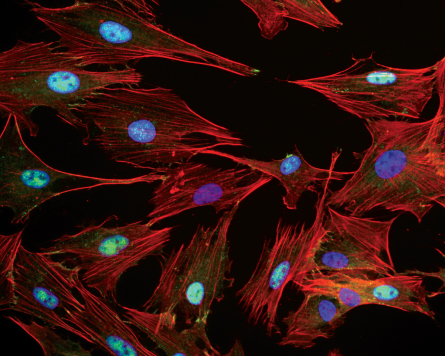
Scientists have found ways to get around this blind spot. When molecules and proteins are labeled with fluorescent dyes, the components inside a cell appear as bright colored dots against the background of a cell’s gel-like filling. Such advances have allowed researchers to see cellular processes unfolding at nanometer scales. Fluorescent probes have also brought to light the wide variation that exists within a population of seemingly alike members.
When sorting through a population of cloned lung cancer cells loaded up with fluorescent probes, systems biologist Steven Altschuler noticed that each cell carried its own combination of the molecules that send signals between cells.
“It was a surprise to us,” says Altschuler, of the University of Texas Southwestern Medical Center at Dallas, who jointly runs a lab with pharmacologist Lani Wu. “It even made us wonder what the definition of a clone was.”
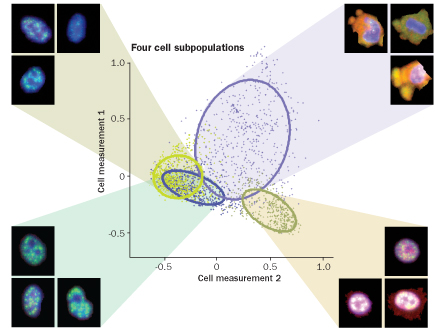
The team developed a way to combine fluorescent microscopy with a type of software designed for face recognition to look for patterns among the cells. The team first labeled the cells by attaching different fluorescent tags to proteins involved in cell signaling. Computers then analyzed pictures of the cells, pixel by pixel, looking for variation in the location and intensity of the molecules. The scientists also trained the computers to identify combinations of markers that commonly appear together — discovering the cellular equivalent of a “blue-eyed blond.”
Using this method, the scientists were able to identify small subpopulations of cells, or stereotypes, based on which molecules appeared and where. Exposing the stereotypes to the anticancer drug Taxol revealed that different subpopulations respond differently to the drug. In May 2010 in Molecular Systems Biology, the team reported on how individual cell differences could help predict the outcome of treatment.
“That’s definitely where we would like to go,” Altschuler says. “The dream is that you would be able to look at a small number of cells from a cancer tumor and try to predict the optimal combination of drugs to treat that particular tumor.”
Though doctors currently use various profiling techniques to determine the best treatment, Altschuler says this new approach differs because, in principle, it would allow a doctor to look at every cell in a tumor biopsy, identifying clues that would get lost in a “population averaging” approach.
Multiplying traits
But fluorescent microscopy has its own limitations. Scientists can look for only a handful of variations, or biomarkers, in a cell at one time. Though cells carrying biomarker A could be distinguished from those carrying B or C, for example, scientists can’t simultaneously scan the cells for a second characteristic that might be important in understanding the first.
Altschuler and Wu’s team has found ways to maximize the use of fluorescent probes to get more details on cell-to-cell differences. Using three out of four available probes to home in on subpopulations of interest, a fourth probe can be free to search for a second variation within each of those groups. By running a series of experiments, rotating the fourth marker’s target each time, scientists can get more detailed information on members in each group.
Say you’re trying to understand people at a baseball stadium, instead of cells, and you want to figure out differences in the way the players, vendors and fans dress. Three out of four fluorescent probes would be used to separate people into each of these groups. The fourth probe could then search for a specific item, such as sneakers, which may be worn by members in each group. In subsequent experiments, the first three probes remain set to identify players, vendors and fans, while the fourth spots people sporting home-team jerseys, then baseball caps and so on. A computer program stitches together the information gleaned from the serial studies to show what people in each group wear.
In this process, a scientist will never look at the jerseys and caps on any given person at the game, because only one measurement can be made at once. “Still, reliable information can be gained,” Altschuler says. His group is now profiling cells in this way.
At her University of North Carolina lab, Allbritton and colleagues are developing ways to study multiple variables beyond the limitations of microscopy. By employing what they call “separation techniques,” the researchers have the potential to measure the activity of many enzymes — a dozen or so — at once. Each cell gets its own well in a grid containing thousands of rows and columns.
Cells are first loaded with various enzyme substrates, substances that work with an enzyme to produce a specific reaction. The cells are then killed and their contents released. By separating out each of the chemicals that were put into the cells as well as the products the cells make, scientists get a readout on what’s going on inside.
Because each cell sits in its own well, scientists can analyze the cells separately. Comparing the reactions of different cells is key, Allbritton says, because a study of a single cell won’t reveal much variability.
“That’s like looking at one human and trying to understand how variable people are,” she says. “You would assume if you got a redheaded person that everyone had red hair — and was 5-foot-4 and was female.”
Allbritton is using this technique to measure protein activity in human cells. Her studies currently focus on kinases, proteins that play a vital role in cell signaling and the regulation of gene activity.
Live on and inform
Such clues to what’s going on inside single cells often come at a price: Cells are damaged or killed in the process, providing only a single snapshot in time. This makes it hard to predict how a single cell will respond to a given stimulus.
J. Christopher Love, a chemical engineer-turned immunologist at MIT, combines the idea of separating cells and the helpfulness of fluorescent dyes to follow biochemical processes over time in living cells. His lab takes an array of subnanoliter-sized wells and fills each with either a single immune cell or a small group of them. With tens of thousands of wells, the unit looks like an ice cube tray the size of a microscope slide. Fluorescent markers identify the various proteins in or on each cell, distinguishing one cell type from another.
Over the course of several hours or days, Love studies the proteins and chemicals secreted by the cells to see how they change. Borrowing a technique from the printing of U.S. dollar bills, he presses the microarray against a glass plate to get a printout (so to speak) of the proteins and chemicals secreted by the cells. Unlike Allbritton’s work, the method doesn’t kill the cells because information is captured from the outside, on the glass plate.
Love is applying this interrogation method to ask questions that are relevant to chronic diseases such as HIV, autoimmunity, cancer and allergies. One ongoing study tracks how allergic patients become desensitized to milk allergens, measuring how the patients’ white blood cells respond to milk extracts. The study looks at how many cells are responding, what kinds of immune proteins are being produced and how much of each protein is produced by each cell.
By analyzing large numbers of single events and interactions between small groups of cells in concert, Love can spot rare types of cells and events that are linked to disease. Certain T cells and B cells found in diseases like multiple sclerosis or diabetes, for example, typically are present at rates of 1 in 10,000 to 1 in 100,000 of the circulating T cells in a blood sample, he says. Without looking at many, many cells, he wouldn’t be able to spot or study the interactions of these rare types.
“Single-cell analysis is a bit of a misnomer in the sense that you’re not really looking at one cell,” Love says. “You’re resolving to one cell, but you are needing to look at thousands, if not tens or hundreds of thousands, of cells to understand what that individual measurement looks like in relation to the rest of the distribution.”
Single microbes
Studies of cell-to-cell differences also promise to reveal new information about single-celled organisms. For a long time, biologists thought that members of a microbe colony were all basically the same, and that those cloned from a single cell, because they were genetically identical, should react to environmental challenges in the very same way.
Scientists now know that individual traits may, in fact, serve as drivers to boost the robustness and resistance of microbial populations, says Michael Konopka, a biochemical engineer at the University of Washington in Seattle. In the October 2010 issue of Nature Chemical Biology, he and coauthor Mary Lidstrom outlined how some members of a microbial colony will stop growing in times of stress, moving into a dormant state that allows them to ride out the bad conditions while their neighbors perish.
Understanding how individuals adapt to environmental conditions, such as temperature fluctuations, could help engineers find ways to boost certain types of chemical processes used in manufacturing, Konopka says. Some naturally occurring microbes, for example, may have a built-in capacity to do the chemical reactions that industrial researchers want to perform.
Others research teams are developing ways to determine the complete genetic blueprint of single-celled organisms to get insight into whole populations of cells that are otherwise difficult to study. Stanford University scientists working under the direction of Stephen Quake are plucking single cells from populations found in pond scum, soil or even plaque on teeth, to ask a seemingly simple question: Who are you?
Conventional genetic analysis techniques require a large, pure sample of the microbes, says Paul Blainey, a postdoctoral researcher in Quake’s lab. “That usually requires that you’re able to isolate the particular organism and grow up a whole huge beaker of it in the lab.” Unfortunately, the chemical reaction used to amplify tiny amounts of DNA is notoriously prone to contamination.
To get around this problem, the Stanford group is using a laser tweezer to sort individual microbes inside tiny, automated devices designed to analyze minute traces of DNA. Enclosed within this miniature lab, microbes and their DNA are safe from contaminants.
In 2007, Quake’s group analyzed the genome of a single bacterium found in dental plaque taken from a person’s mouth. This year the team turned to a less familiar class of single-celled organisms, the Archaea: Online February 22 in PLoS One, the researchers published findings on the genome of a single-celled ammonia-eater from San Francisco Bay.
Genetic information gleaned from an individual microbe, combined with findings on cell-to-cell differences in behavior, may provide new insights into the thousands of unknown or barely known populations found in water and soil. Such insight could in turn offer clues to how microbes act as infectious agents and how they develop resistance to antibiotics.
Despite recent gains in studying single cells and how they differ, there’s much work to be done, Allbritton says. Current techniques still allow only a small number — fewer than two dozen — of the thousands of molecular components that float around in cells to be measured at one time.
“It’s embarrassing,” she says. “We’ve made a lot of progress, but you can see how far we have to go.”
With such a small subset of cellular components under study, scientists’ perception of how cells operate may be colored by what they see, she adds. The big push now is to develop ways to measure more of the cell’s contents, and see how they change over time and work in relationship to each other.
“If we can see everything,” Allbritton says, “it might dramatically change how we view the single cell.”